Compound details
Deferoxamine
| Compound ID | CDAMM00787 |
|---|---|
| Common name | Deferoxamine | IUPAC name | N-[5-[[4-[5-[acetyl(hydroxy)amino]pentylamino]-4-oxobutanoyl]-hydroxyamino]pentyl]-N\'-(5-aminopentyl)-N\'-hydroxybutanediamide |
| Molecular formula | C25H48N6O8 |
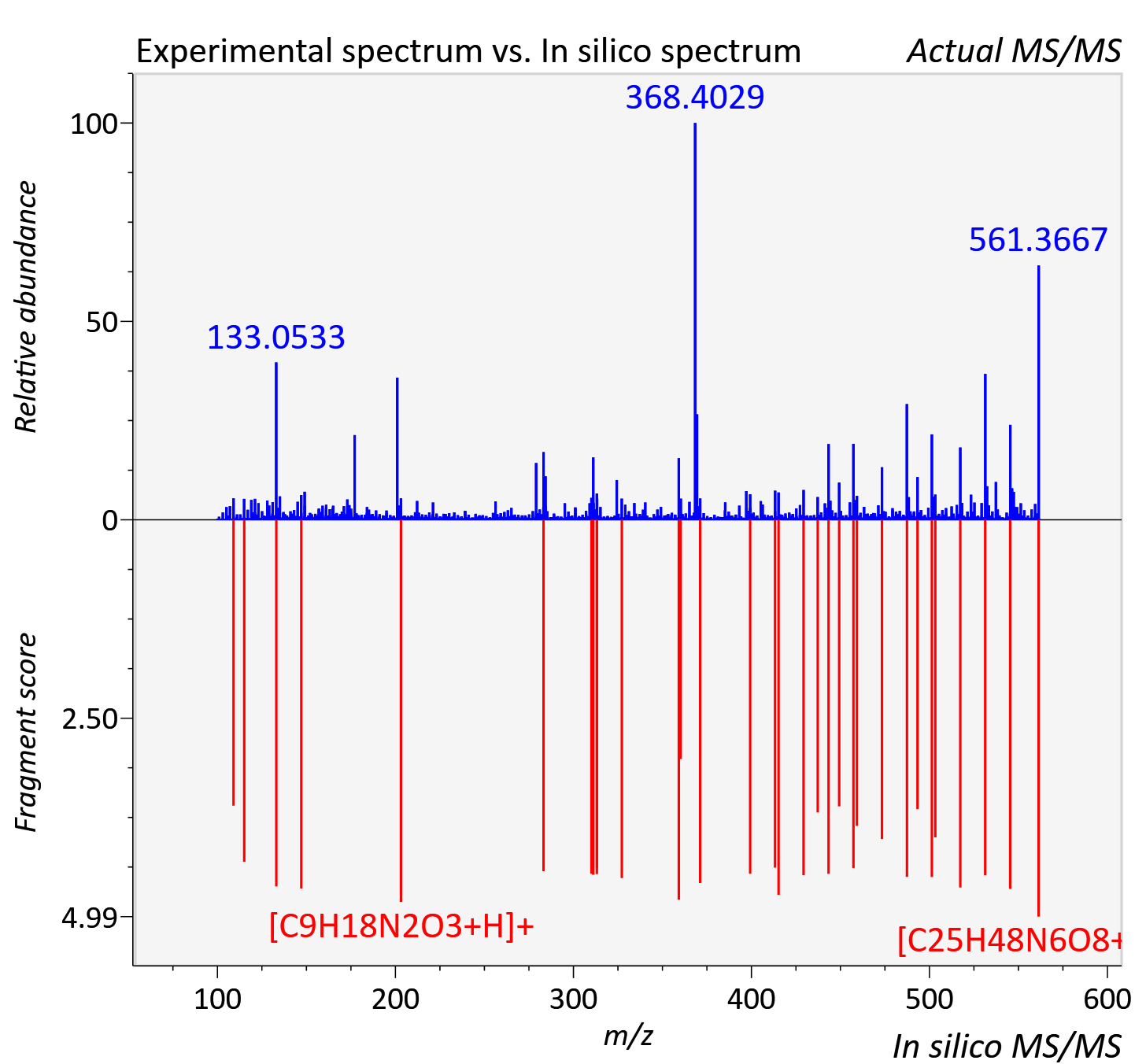
Experimental data
| Retention time | 13.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 561.367 | Theoretical mz | 561.36 |
| Error | 11.98 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.0557 |
Identifiers and class information
| Inchi key | UBQYURCVBFRUQT-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C(NCCCCCN(O)C(=O)C)CCC(=O)N(O)CCCCCNC(=O)CCC(=O)N(O)CCCCCN |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q9H3R0 | KDM4C | Lysine-specific demethylase 4C | T73863 | SEA |
| O75164 | KDM4A | Lysine-specific demethylase 4A | T18904 | SEA |
| Q6QHF9 | PAOX | Polyamine oxidase | T88699 | SEA |
| Q6ZMT4 | KDM7A | Lysine-specific demethylase 7A/7B | T18784 | SEA |
| P42263 | GRIA3 | Glutamate receptor AMPA 3 | T62391 | SEA |
