Compound details
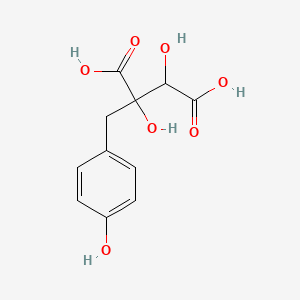
(2R,3S)-Piscidic acid
| Compound ID | CDAMM00724 |
|---|---|
| Common name | (2R,3S)-Piscidic acid | IUPAC name | 2,3-dihydroxy-2-[(4-hydroxyphenyl)methyl]butanedioic acid |
| Molecular formula | C11H12O7 |
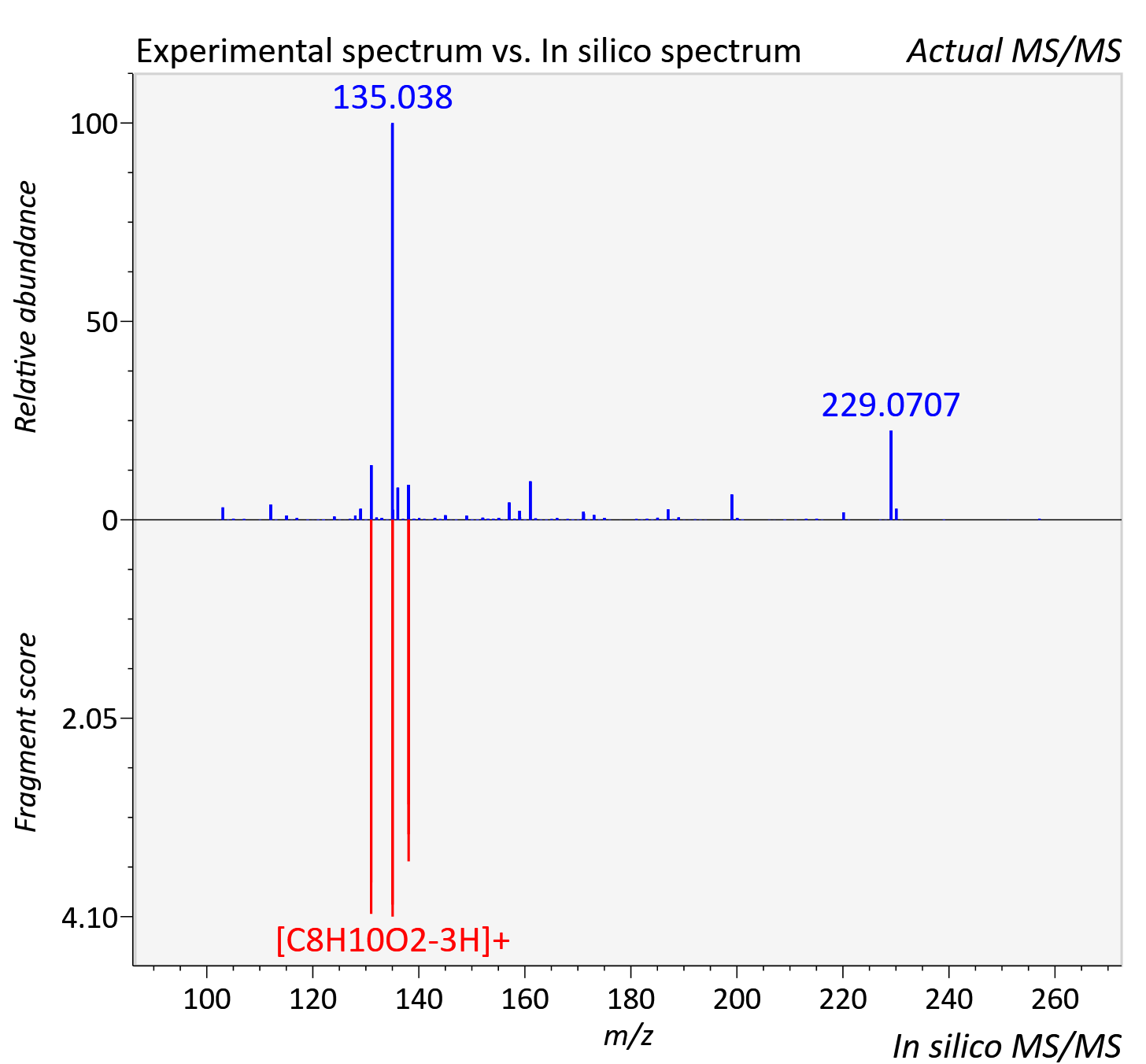
Experimental data
| Retention time | 9.64 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 257.065 | Theoretical mz | 257.065 |
| Error | 0.09 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.9543 |
Identifiers and class information
| Inchi key | TUODPMGCCJSJRH-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(O)C(O)C(O)(C(=O)O)CC1=CC=C(O)C=C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Phenylpropanoic acids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08473 | MME | Neprilysin | T05409 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| P80404 | ABAT | Gamma-amino-N-butyrate transaminase | T59045 | SEA |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| P07101 | TH | Tyrosine 3-hydroxylase | T62390 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| Q6P179 | ERAP2 | Endoplasmic reticulum aminopeptidase 2 | T65818 | SEA |
| Q86YT5 | SLC13A5 | Solute carrier family 13 member 5 | T81547 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| P53396 | ACLY | ATP-citrate synthase | T52189 | SEA |
| P12004 | PCNA | Proliferating cell nuclear antigen | T21782 | SEA |
| P15169 | CPN1 | Carboxypeptidase N, catalytic subunit | T34471 | SEA |
| P45974 | USP5 | Ubiquitin carboxyl-terminal hydrolase 5 | T17067 | SEA |
| P51649 | ALDH5A1 | Succinate semialdehyde dehydrogenase | T13259 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T05409 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08473 | MME |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T59045 | DI0060 | Brain cancer | [ICD-11: 2A00] | P80404 | ABAT |
| T59045 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P80404 | ABAT |
| T59045 | DI0135 | Epileptic encephalopathy | [ICD-11: 8A62] | P80404 | ABAT |
| T59045 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P80404 | ABAT |
| T59045 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P80404 | ABAT |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T62390 | DI0019 | Adrenomedullary hyperfunction | [ICD-11: 5A75] | P07101 | TH |
| T62390 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P07101 | TH |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T52189 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | P53396 | ACLY |
| T21782 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P12004 | PCNA |
| T17067 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P45974 | USP5 |
| T17067 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P45974 | USP5 |
| T13259 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P51649 | ALDH5A1 |
| T13259 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P51649 | ALDH5A1 |
