Compound details
Pipermethystine
| Compound ID | CDAMM00712 |
|---|---|
| Common name | Pipermethystine | IUPAC name | [6-oxo-1-(3-phenylpropanoyl)-2,3-dihydropyridin-3-yl] acetate |
| Molecular formula | C16H17NO4 |
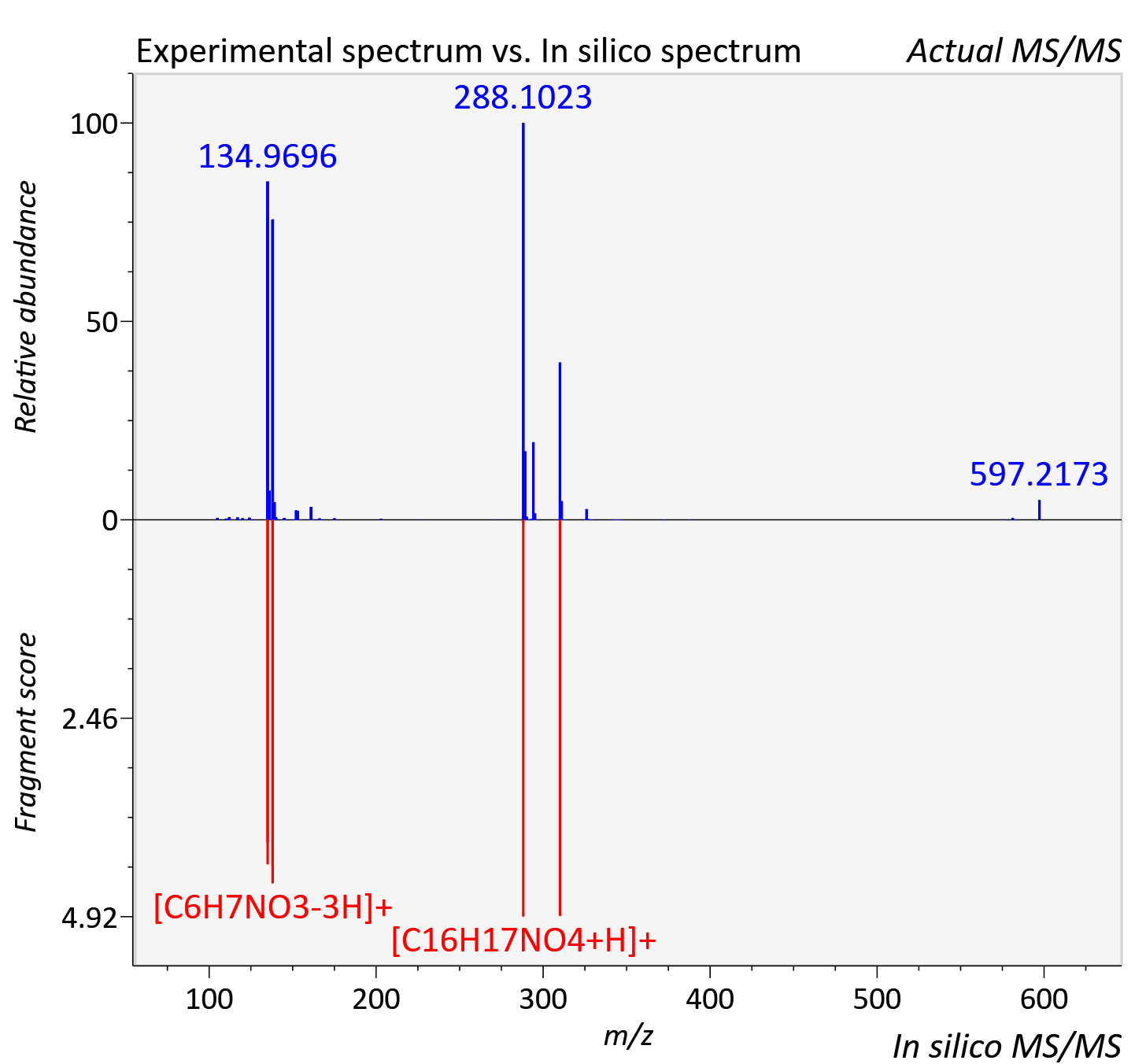
Experimental data
| Retention time | 7.4 |
|---|---|
| Adduct | [2M+Na]+ |
| Actual mz | 597.218 | Theoretical mz | 597.221 |
| Error | 6.23 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.298 |
Identifiers and class information
| Inchi key | JLNNQCUATONMIT-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(OC1C=CC(=O)N(C(=O)CCC=2C=CC=CC2)C1)C |
| Superclass | Organoheterocyclic compounds |
| Class | Pyridines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SEA |
| Q96RJ0 | TAAR1 | Trace amine-associated receptor 1 (by homology) | T99524 | SEA |
| P32297 | CHRNA3 | Neuronal acetylcholine receptor; alpha3/beta4 | T74166 | SEA |
| P42330 | AKR1C3 | Aldo-keto-reductase family 1 member C3 | T60857 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| O14842 | FFAR1 | Free fatty acid receptor 1 | T25608 | SEA |
| P08183 | ABCB1 | P-glycoprotein 1 | T25258 | SEA |
| P62942 | FKBP1A | FK506-binding protein 1A | T76059 | SEA |
| P48147 | PREP | Prolyl endopeptidase | T86161 | SEA |
| Q99685 | MGLL | Monoglyceride lipase | T18664 | SEA |
| P41229 | KDM5C | Lysine-specific demethylase 5C | T85540 | SEA |
| Q9Y2K7 | KDM2A | Lysine-specific demethylase 2A | T71949 | SEA |
| P42574 | CASP3 | Caspase-3 | T57943 | SEA |
| P09601 | HMOX1 | Heme oxygenase 1 (by homology) | T25703 | SEA |
| Q9Y271 | CYSLTR1 | Cysteinyl leukotriene receptor 1 | T71192 | SEA |
| P30926 | CHRNB4 | Neuronal acetylcholine receptor; alpha2/beta4 | T73724 | SEA |
| O43895 | XPNPEP2 | Xaa-Pro aminopeptidase 2 | T68320 | SEA |
| Q9NS75 | CYSLTR2 | Cysteinyl leukotriene receptor 2 | T74238 | SEA |
| P17787 | CHRNB2 | Nicotinic acetylcholine receptor alpha2/beta2 | T82543 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T99524 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | Q96RJ0 | TAAR1 |
| T99524 | DI0149 | Focal/segmental autonomic disorder | [ICD-11: 8D8A] | Q96RJ0 | TAAR1 |
| T99524 | DI0290 | Narcolepsy | [ICD-11: 7A20] | Q96RJ0 | TAAR1 |
| T74166 | DI0105 | Cough | [ICD-11: MD12] | P32297 | CHRNA3 |
| T60857 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P42330 | AKR1C3 |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T25608 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | O14842 | FFAR1 |
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T76059 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P62942 | FKBP1A |
| T76059 | DI0349 | Pruritus | [ICD-11: EC90] | P62942 | FKBP1A |
| T76059 | DI0413 | Transplant rejection | [ICD-11: NE84] | P62942 | FKBP1A |
| T86161 | DI0092 | Coeliac disease | [ICD-11: DA95] | P48147 | PREP |
| T86161 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | P48147 | PREP |
| T86161 | DI0238 | Lung cancer | [ICD-11: 2C25] | P48147 | PREP |
| T86161 | DI0265 | Mild neurocognitive disorder | [ICD-11: 6D71] | P48147 | PREP |
| T18664 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q99685 | MGLL |
| T18664 | DI0409 | Tic disorder | [ICD-11: 8A05] | Q99685 | MGLL |
| T57943 | DI0060 | Brain cancer | [ICD-11: 2A00] | P42574 | CASP3 |
| T25703 | DI0294 | Neonatal hyperbilirubinaemia | [ICD-11: KA87] | P09601 | HMOX1 |
| T71192 | DI0037 | Asthma | [ICD-11: CA23] | Q9Y271 | CYSLTR1 |
| T73724 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P30926 | CHRNB4 |
| T74238 | DI0037 | Asthma | [ICD-11: CA23] | Q9NS75 | CYSLTR2 |
| T82543 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P17787 | CHRNB2 |
