Compound details
Yunngnin A
| Compound ID | CDAMM00673 |
|---|---|
| Common name | Yunngnin A | IUPAC name | 7-hydroxy-5,6-dimethoxy-2-oxochromene-8-carbaldehyde |
| Molecular formula | C12H10O6 |
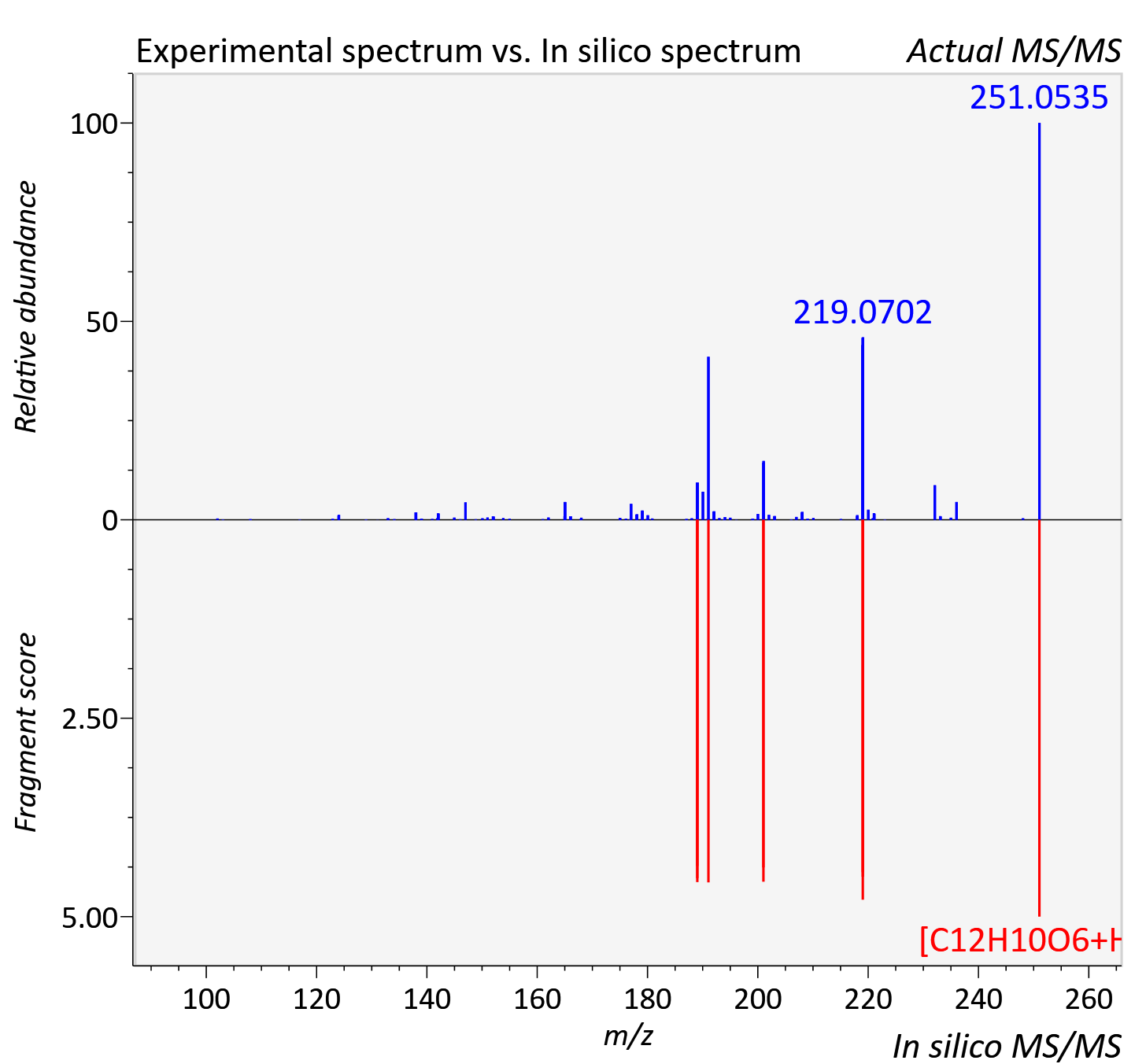
Experimental data
| Retention time | 3.36 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 251.054 | Theoretical mz | 251.055 |
| Error | 5.88 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2485 |
Identifiers and class information
| Inchi key | IVKYXTYKRIADDE-UHFFFAOYSA-N |
|---|---|
| Smiles | O=CC1=C(O)C(OC)=C(OC)C=2C=CC(=O)OC12 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Coumarins and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SEA |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| Q08499 | PDE4D | Phosphodiesterase 4D | T02001 | SEA |
| O14980 | XPO1 | Exportin-1 | T51407 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| Q9HC98 | NEK6 | Serine/threonine-protein kinase NEK6 | T78992 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| P02768 | ALB | Serum albumin | T63068 | SEA |
| P16220 | CREB1 | Cyclic AMP-responsive element-binding protein | T92098 | SEA |
| Q09470 | KCNA1 | Voltage-gated potassium channel Kv1.1 | T85272 | SEA |
| P22459 | KCNA4 | Voltage-gated A-type potassium channel | T15665 | SEA |
| P16389 | KCNA2 | Voltage-gated potassium channel Kv1.2 | T19492 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T02001 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q08499 | PDE4D |
| T51407 | DI0234 | Liposarcoma | [ICD-11: 2B59] | O14980 | XPO1 |
| T51407 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | O14980 | XPO1 |
| T51407 | DI0395 | Stomach cancer | [ICD-11: 2B72] | O14980 | XPO1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T63068 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P02768 | ALB |
| T63068 | DI0059 | Bowel habit change | [ICD-11: ME05] | P02768 | ALB |
| T63068 | DI0077 | Cholelithiasis | [ICD-11: DC11] | P02768 | ALB |
| T63068 | DI0202 | Inborn energy metabolism error | [ICD-11: 5C53] | P02768 | ALB |
| T63068 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P02768 | ALB |
| T63068 | DI0343 | Preprocedural examination | [ICD-11: QA0B] | P02768 | ALB |
| T63068 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P02768 | ALB |
| T63068 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P02768 | ALB |
| T63068 | DI0420 | Unspecific body region injury | [ICD-11: ND56] | P02768 | ALB |
