Compound details
(S)-Isowillardiine
| Compound ID | CDAMM00664 |
|---|---|
| Common name | (S)-Isowillardiine | IUPAC name | 2-amino-3-(2,4-dioxo-1H-pyrimidin-3-yl)propanoic acid |
| Molecular formula | C7H9N3O4 |
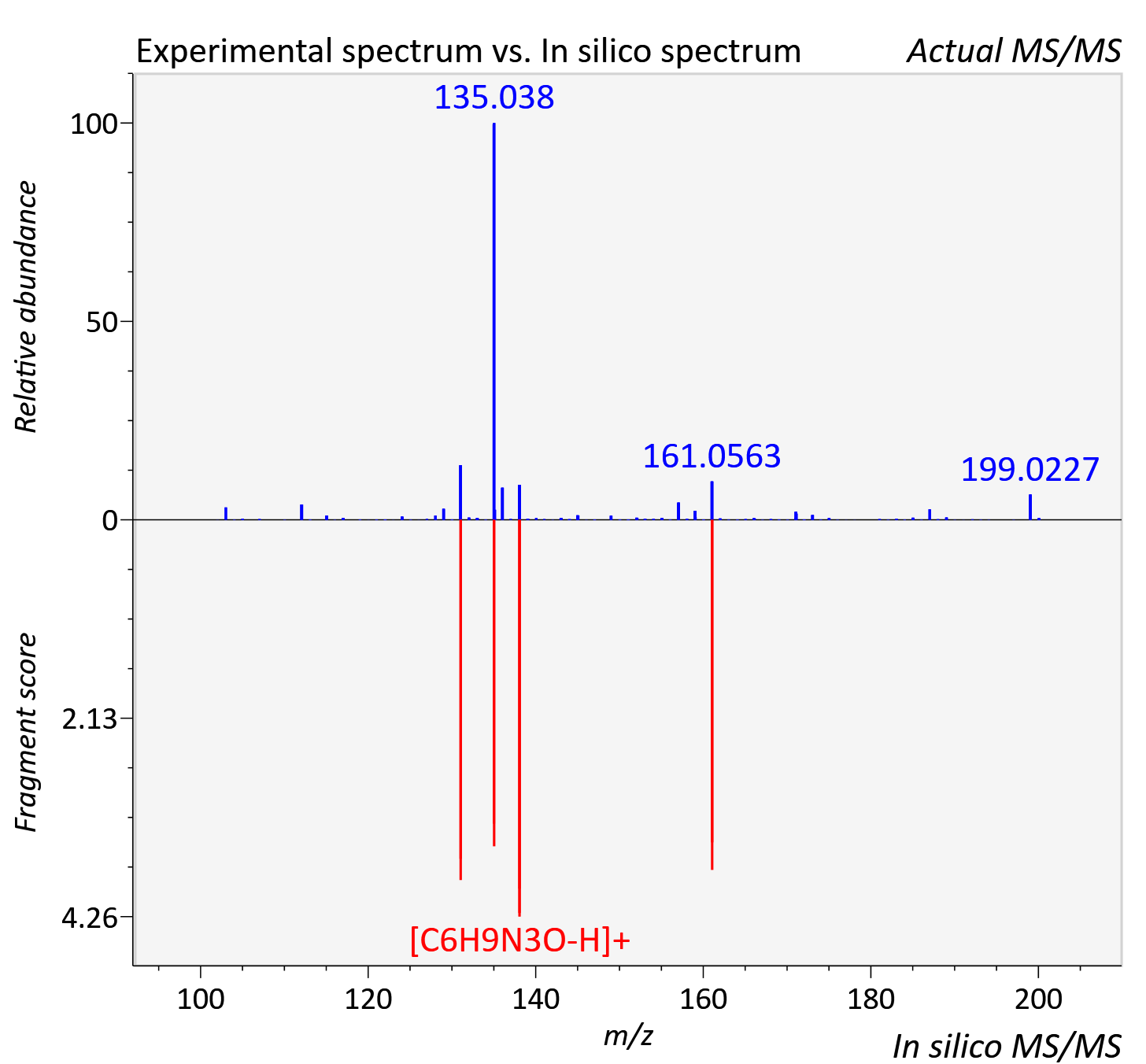
Experimental data
| Retention time | 9.64 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 200.066 | Theoretical mz | 200.066 |
| Error | 0.88 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.629 |
Identifiers and class information
| Inchi key | AZSWUZQIIMMKOZ-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C(O)C(N)CN1C(=O)C=CN=C1O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P43005 | SLC1A1 | Excitatory amino acid transporter 3 | T31721 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SwissTargetPrediction and SEA |
| P48058 | GRIA4 | Glutamate receptor ionotropic, AMPA 4 | T96235 | SwissTargetPrediction |
| Q16478 | GRIK5 | Glutamate receptor ionotropic kainate 5 | T87930 | SwissTargetPrediction and SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| P42262 | GRIA2 | Glutamate receptor ionotropic, AMPA 2 | T42392 | SwissTargetPrediction and SEA |
| P07101 | TH | Tyrosine 3-hydroxylase | T62390 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SEA |
| Q01650 | SLC7A5 | L-type amino acid transporter 1 (by homology) | T48330 | SEA |
| P43003 | SLC1A3 | Excitatory amino acid transporter 1 | T86582 | SEA |
| P05089 | ARG1 | Arginase-1 (by homology) | T63224 | SEA |
| P12004 | PCNA | Proliferating cell nuclear antigen | T21782 | SEA |
| Q9UBC3 | DNMT3B | DNA (cytosine-5)-methyltransferase 3B | T65501 | SEA |
| Q9UPY5 | SLC7A11 | Cystine/glutamate transporter | T11615 | SEA |
| P42263 | GRIA3 | Glutamate receptor AMPA 3 | T62391 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T96235 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P48058 | GRIA4 |
| T42392 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42262 | GRIA2 |
| T62390 | DI0019 | Adrenomedullary hyperfunction | [ICD-11: 5A75] | P07101 | TH |
| T62390 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P07101 | TH |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T63224 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P05089 | ARG1 |
| T21782 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P12004 | PCNA |
| T65501 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9UBC3 | DNMT3B |
| T11615 | DI0054 | Body-focused behaviour disorder | [ICD-11: 6B25] | Q9UPY5 | SLC7A11 |
| T62391 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42263 | GRIA3 |
