Compound details
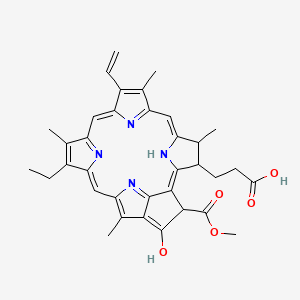
13-epi-Phaeophorbide a
| Compound ID | CDAMM00649 |
|---|---|
| Common name | 13-epi-Phaeophorbide a | IUPAC name | 3-(16-ethenyl-11-ethyl-4-hydroxy-3-methoxycarbonyl-12,17,21,26-tetramethyl-7,23,24,25-tetrazahexacyclo[18.2.1.15,8.110,13.115,18.02,6]hexacosa-1,4,6,8(26),9,11,13(25),14,16,18(24),19-undecaen-22-yl)propanoic acid |
| Molecular formula | C35H36N4O5 |
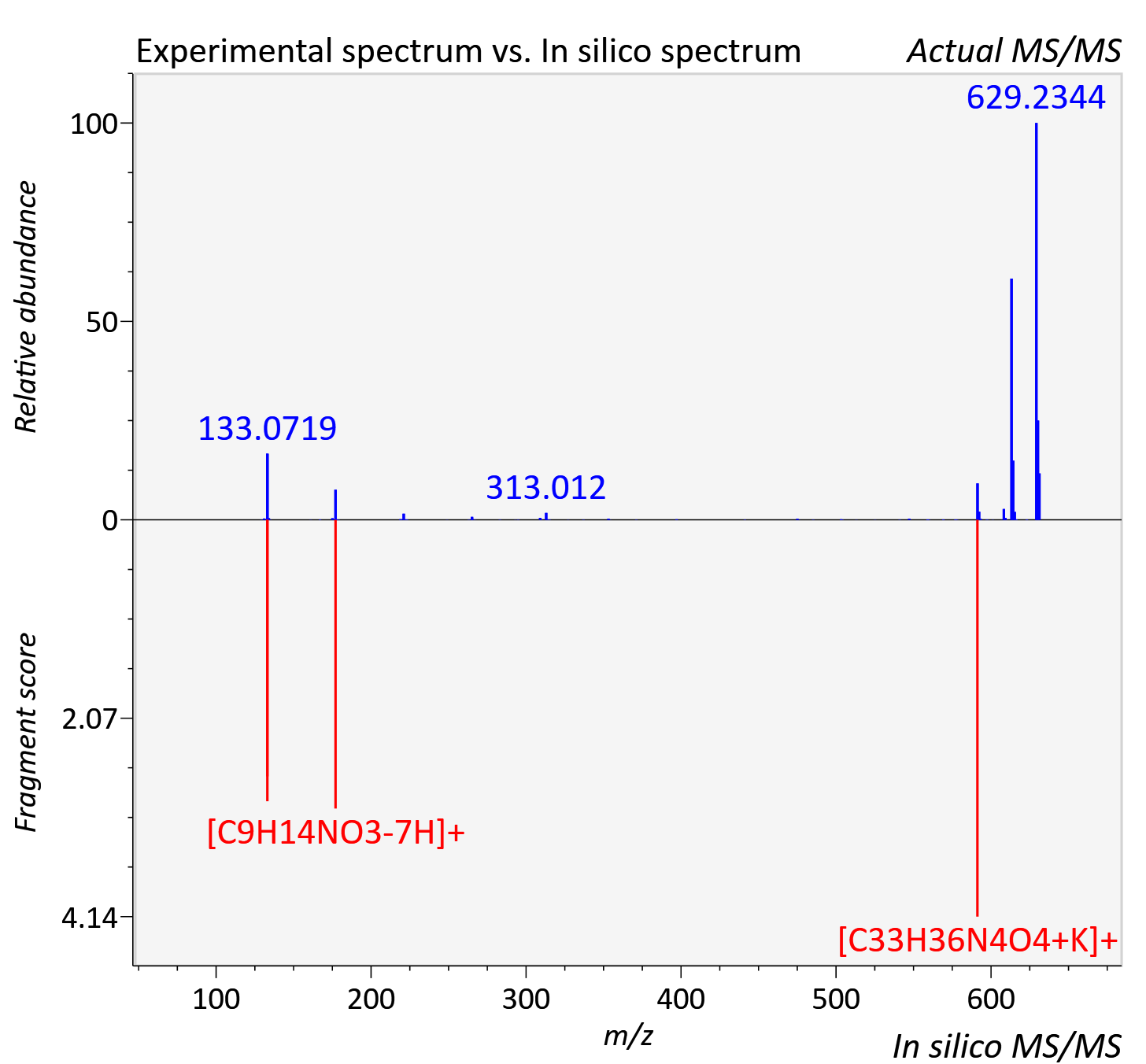
Experimental data
| Retention time | 0.51 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 631.237 | Theoretical mz | 631.232 |
| Error | 6.88 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.4696 |
Identifiers and class information
| Inchi key | NSFSLUUZQIAOOX-DBPJEZTQNA-N |
|---|---|
| Smiles | O=C(O)CCC1C2=NC(=CC=3NC(C=C4N=C(C=C5NC=6C(C(=O)C(C(=O)OC)C62)=C5C)C(=C4C)CC)=C(C=C)C3C)C1C |
| Superclass | Organoheterocyclic compounds |
| Class | Tetrapyrroles and derivatives |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T75440 | DI0033 | Anxiety disorder | [ICD-11: 6B00-6B0Z] | P30536 | TSPO |
| T75440 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P30536 | TSPO |
| T75440 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P30536 | TSPO |
| T75440 | DI0216 | Intentional self-harm | [ICD-11: PC91] | P30536 | TSPO |
| T75440 | DI0271 | Mood/affect symptom | [ICD-11: MB24] | P30536 | TSPO |
| T75440 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P30536 | TSPO |
