Compound details
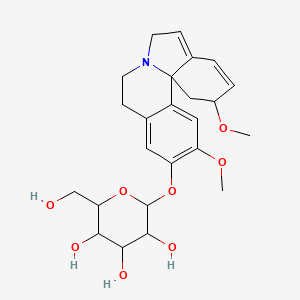
Glucoerysodine
| Compound ID | CDAMM00619 |
|---|---|
| Common name | Glucoerysodine | IUPAC name | 2-[(2,12-dimethoxy-2,6,8,9-tetrahydro-1H-indolo[7a,1-a]isoquinolin-11-yl)oxy]-6-(hydroxymethyl)oxane-3,4,5-triol |
| Molecular formula | C24H31NO8 |
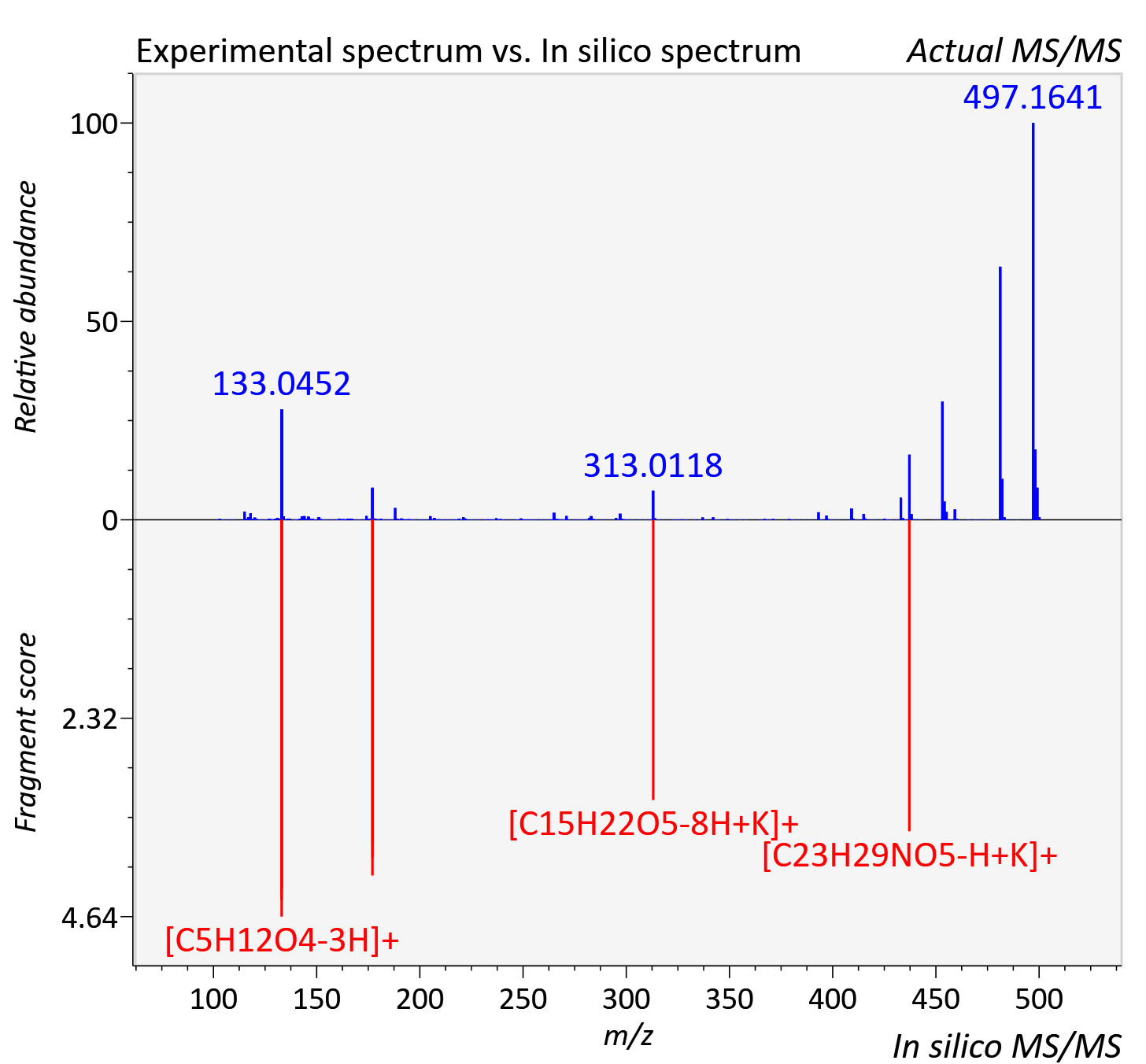
Experimental data
| Retention time | 0.42 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 500.168 | Theoretical mz | 500.168 |
| Error | 0.72 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7307 |
Identifiers and class information
| Inchi key | LDKVUIURMJHFPP-WDHHSZOHNA-N |
|---|---|
| Smiles | OCC1OC(OC=2C=C3C(=CC2OC)C45C(C=CC(OC)C4)=CCN5CC3)C(O)C(O)C1O |
| Superclass | Alkaloids and derivatives |
| Class | Erythrina alkaloids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SwissTargetPrediction |
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SwissTargetPrediction |
| P29274 | ADORA2A | Adenosine A2a receptor | T77365 | SwissTargetPrediction |
| P12821 | ACE | Angiotensin-converting enzyme | T98311 | SwissTargetPrediction |
| P13866 | SLC5A1 | Sodium/glucose cotransporter 1 | T54771 | SwissTargetPrediction |
| P08473 | MME | Neprilysin | T05409 | SwissTargetPrediction |
| P35372 | OPRM1 | Mu opioid receptor | T47768 | SwissTargetPrediction |
| P31639 | SLC5A2 | Sodium/glucose cotransporter 2 | T30085 | SwissTargetPrediction |
| P60568 | IL2 | Interleukin-2 | T61698 | SEA |
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q9NZ08 | ERAP1 | Endoplasmic reticulum aminopeptidase 1 | T72849 | SEA |
| P16444 | DPEP1 | Renal dipeptidase | T41201 | SwissTargetPrediction |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T77365 | DI0319 | Orthostatic hypotension | [ICD-11: BA21] | P29274 | ADORA2A |
| T77365 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P29274 | ADORA2A |
| T77365 | DI0359 | Radionuclide imaging | [ICD-11: N.A.] | P29274 | ADORA2A |
| T98311 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P12821 | ACE |
| T98311 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P12821 | ACE |
| T54771 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P13866 | SLC5A1 |
| T54771 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P13866 | SLC5A1 |
| T54771 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P13866 | SLC5A1 |
| T05409 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08473 | MME |
| T47768 | DI0013 | Acute pain | [ICD-11: MG31] | P35372 | OPRM1 |
| T47768 | DI0059 | Bowel habit change | [ICD-11: ME05] | P35372 | OPRM1 |
| T47768 | DI0087 | Chronic pain | [ICD-11: MG30] | P35372 | OPRM1 |
| T47768 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P35372 | OPRM1 |
| T47768 | DI0105 | Cough | [ICD-11: MD12] | P35372 | OPRM1 |
| T47768 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P35372 | OPRM1 |
| T47768 | DI0124 | Digestive system disease | [ICD-11: DE2Z] | P35372 | OPRM1 |
| T47768 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P35372 | OPRM1 |
| T47768 | DI0228 | Large intestine motility disorder | [ICD-11: DB32] | P35372 | OPRM1 |
| T47768 | DI0317 | Opioid use disorder | [ICD-11: 6C43] | P35372 | OPRM1 |
| T47768 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P35372 | OPRM1 |
| T47768 | DI0349 | Pruritus | [ICD-11: EC90] | P35372 | OPRM1 |
| T47768 | DI0374 | Sensation disturbance | [ICD-11: MB40] | P35372 | OPRM1 |
| T30085 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P31639 | SLC5A2 |
| T30085 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P31639 | SLC5A2 |
| T30085 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P31639 | SLC5A2 |
| T61698 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P60568 | IL2 |
| T61698 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P60568 | IL2 |
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T41201 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P16444 | DPEP1 |
