Compound details
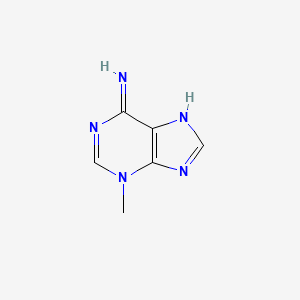
3-Methyladenine
| Compound ID | CDAMM00572 |
|---|---|
| Common name | 3-Methyladenine | IUPAC name | 3-methyl-7H-purin-6-imine |
| Molecular formula | C6H7N5 |
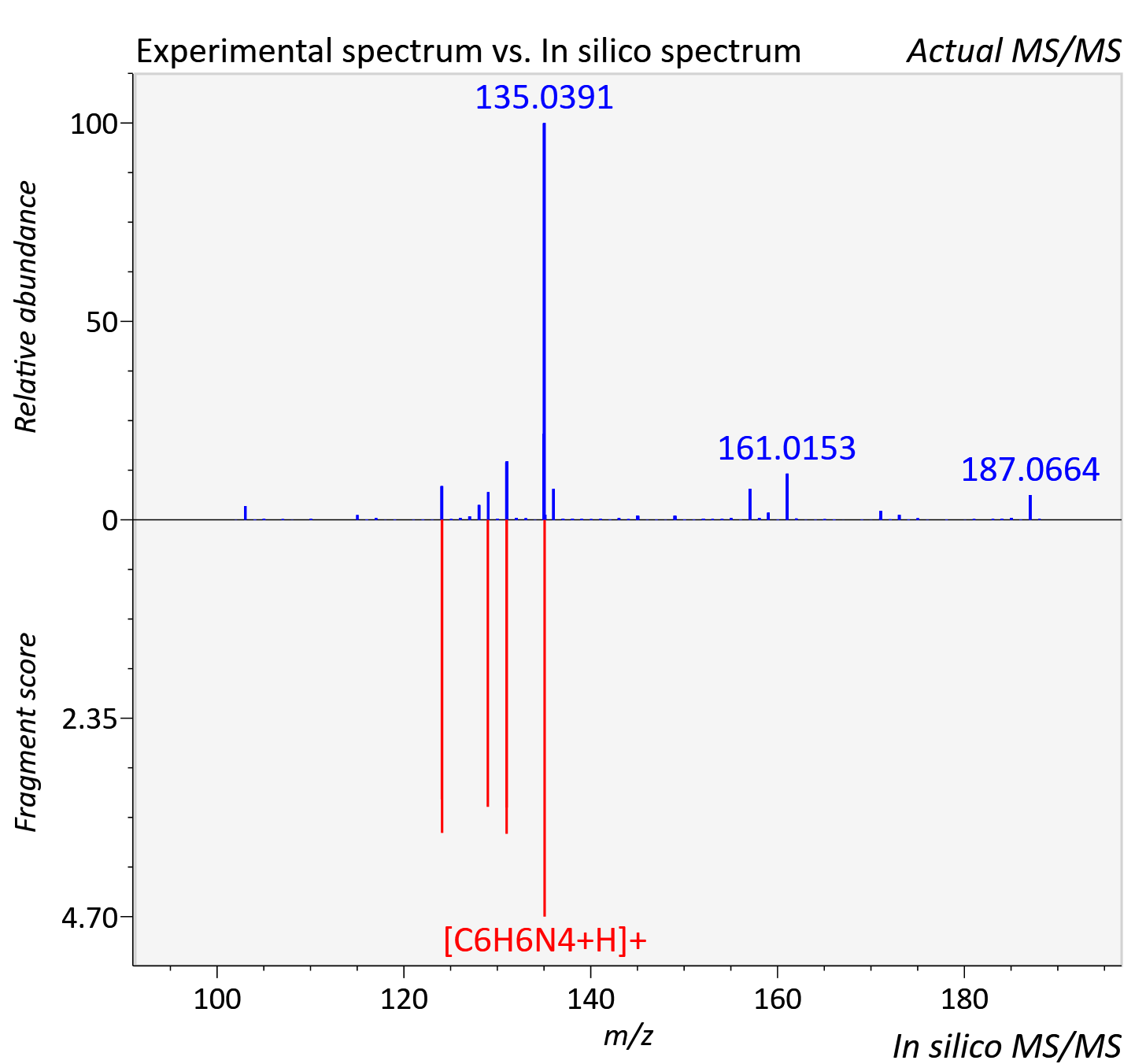
Experimental data
| Retention time | 8.87 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 188.033 | Theoretical mz | 188.033 |
| Error | 2.44 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.3082 |
Identifiers and class information
| Inchi key | FSASIHFSFGAIJM-UHFFFAOYSA-N |
|---|---|
| Smiles | N1=CN=C2C1=C(N=CN2C)N |
| Superclass | Organoheterocyclic compounds |
| Class | Imidazopyrimidines |
