Compound details
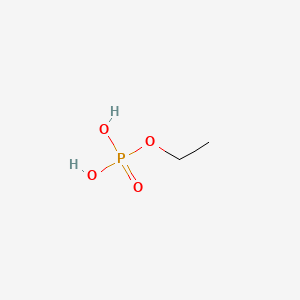
Ethylphosphate
| Compound ID | CDAMM00568 |
|---|---|
| Common name | Ethylphosphate | IUPAC name | ethyl dihydrogen phosphate |
| Molecular formula | C2H7O4P |
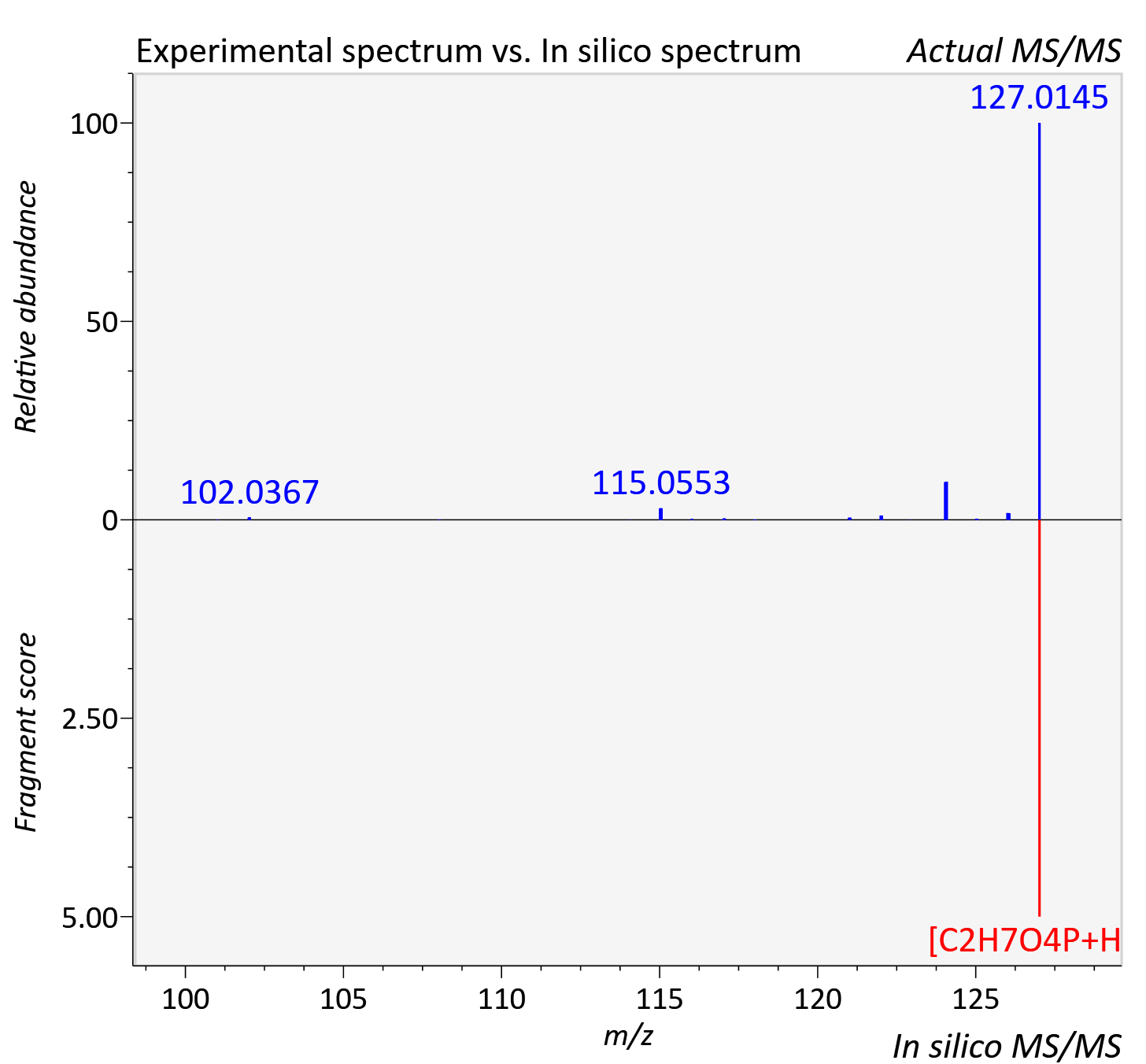
Experimental data
| Retention time | 8.51 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 127.014 | Theoretical mz | 127.015 |
| Error | 7.68 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.3319 |
Identifiers and class information
| Inchi key | ZJXZSIYSNXKHEA-UHFFFAOYSA-N |
|---|---|
| Smiles | O=P(O)(O)OCC |
| Superclass | Organic acids and derivatives |
| Class | Organic phosphoric acids and derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P52209 | PGD | 6-phosphogluconate dehydrogenase | T76497 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| Q99500 | S1PR3 | Sphingosine 1-phosphate receptor Edg-3 | T11241 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9H1C0 | LPAR5 | Lysophosphatidic acid receptor 5 | T18616 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| P51575 | P2RX1 | P2X purinoceptor 1 | T69091 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P53602 | MVD | Diphosphomevalonate decarboxylase | T96862 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
| T11241 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q99500 | S1PR3 |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
