Compound details
Phenol
| Compound ID | CDAMM00565 |
|---|---|
| Common name | Phenol | IUPAC name | phenol |
| Molecular formula | C6H6O |
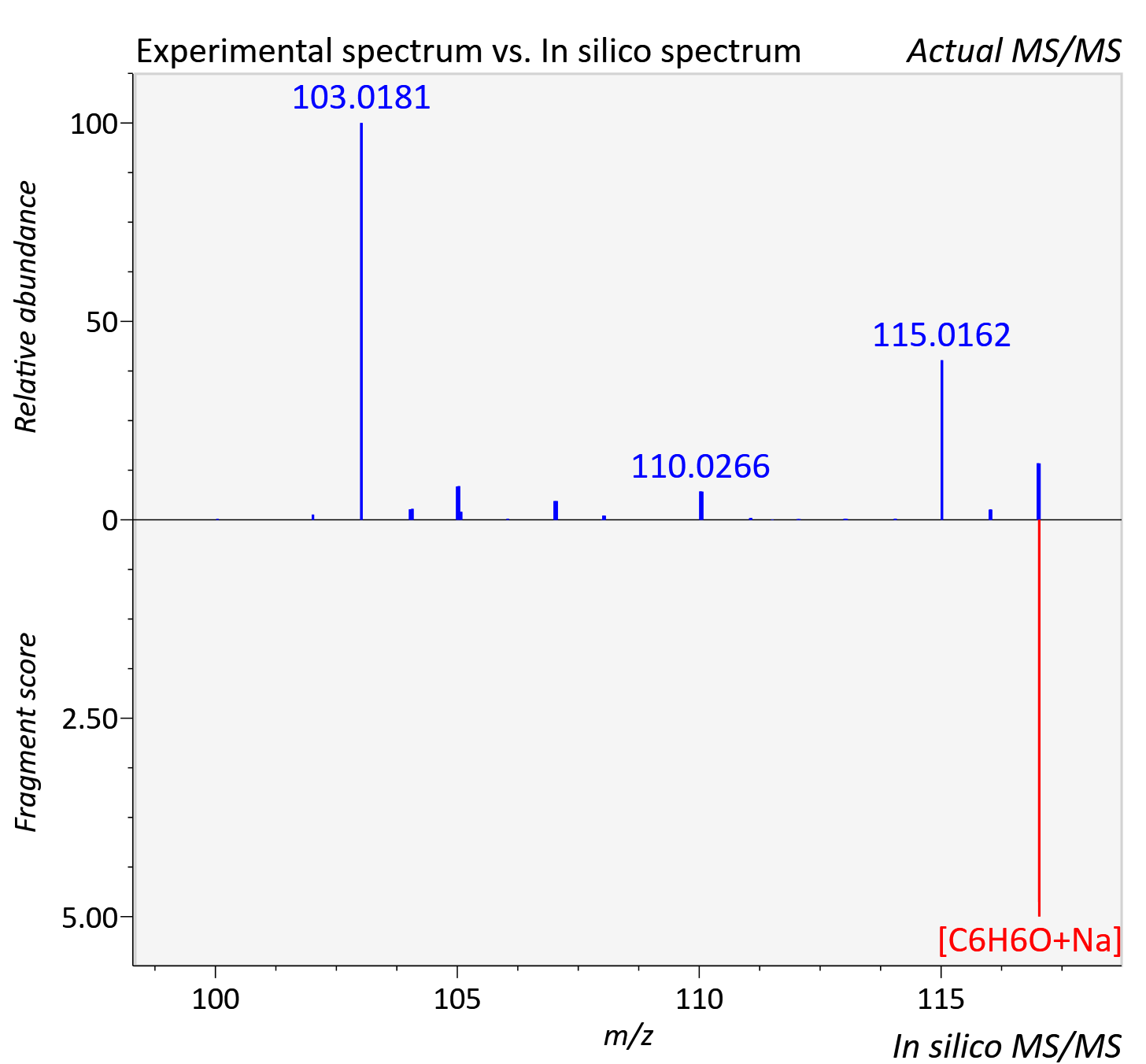
Experimental data
| Retention time | 8.87 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 117.032 | Theoretical mz | 117.031 |
| Error | 8.39 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7913 |
Identifiers and class information
| Inchi key | ISWSIDIOOBJBQZ-UHFFFAOYSA-N |
|---|---|
| Smiles | OC=1C=CC=CC1 |
| Superclass | Benzenoids |
| Class | Phenols |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SwissTargetPrediction and SEA |
| Q16790 | CA9 | Carbonic anhydrase IX | T64567 | SwissTargetPrediction and SEA |
| P02766 | TTR | Transthyretin | T86462 | SEA |
| P00918 | CA2 | Carbonic anhydrase II | T20401 | SwissTargetPrediction and SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SwissTargetPrediction and SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| P80404 | ABAT | Gamma-amino-N-butyrate transaminase | T59045 | SEA |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| P14902 | IDO1 | Indoleamine 2,3-dioxygenase | T89697 | SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| P62508 | ESRRG | Estrogen-related receptor gamma | T59875 | SEA |
| P45974 | USP5 | Ubiquitin carboxyl-terminal hydrolase 5 | T17067 | SEA |
| P51649 | ALDH5A1 | Succinate semialdehyde dehydrogenase | T13259 | SEA |
| P09172 | DBH | Dopamine beta hydroxylase | T74937 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T64567 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q16790 | CA9 |
| T86462 | DI0026 | Amyloidosis | [ICD-11: 5D00] | P02766 | TTR |
| T20401 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P00918 | CA2 |
| T20401 | DI0137 | Essential hypertension | [ICD-11: BA00] | P00918 | CA2 |
| T20401 | DI0166 | Glaucoma | [ICD-11: 9C61] | P00918 | CA2 |
| T20401 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P00918 | CA2 |
| T20401 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P00918 | CA2 |
| T20401 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P00918 | CA2 |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T59045 | DI0060 | Brain cancer | [ICD-11: 2A00] | P80404 | ABAT |
| T59045 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P80404 | ABAT |
| T59045 | DI0135 | Epileptic encephalopathy | [ICD-11: 8A62] | P80404 | ABAT |
| T59045 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P80404 | ABAT |
| T59045 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P80404 | ABAT |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T89697 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P14902 | IDO1 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T17067 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P45974 | USP5 |
| T17067 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P45974 | USP5 |
| T13259 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | P51649 | ALDH5A1 |
| T13259 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P51649 | ALDH5A1 |
| T74937 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P09172 | DBH |
| T74937 | DI0341 | Post-traumatic stress disorder | [ICD-11: 6B40] | P09172 | DBH |
