Compound details
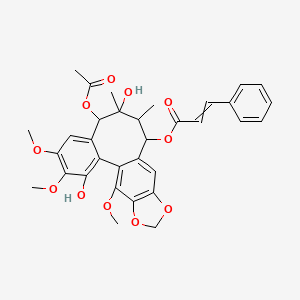
Kadsuphilol L
| Compound ID | CDAMM00548 |
|---|---|
| Common name | Kadsuphilol L | IUPAC name | (8-acetyloxy-3,9-dihydroxy-4,5,19-trimethoxy-9,10-dimethyl-15,17-dioxatetracyclo[10.7.0.02,7.014,18]nonadeca-1(19),2,4,6,12,14(18)-hexaen-11-yl) 3-phenylprop-2-enoate |
| Molecular formula | C33H34O11 |
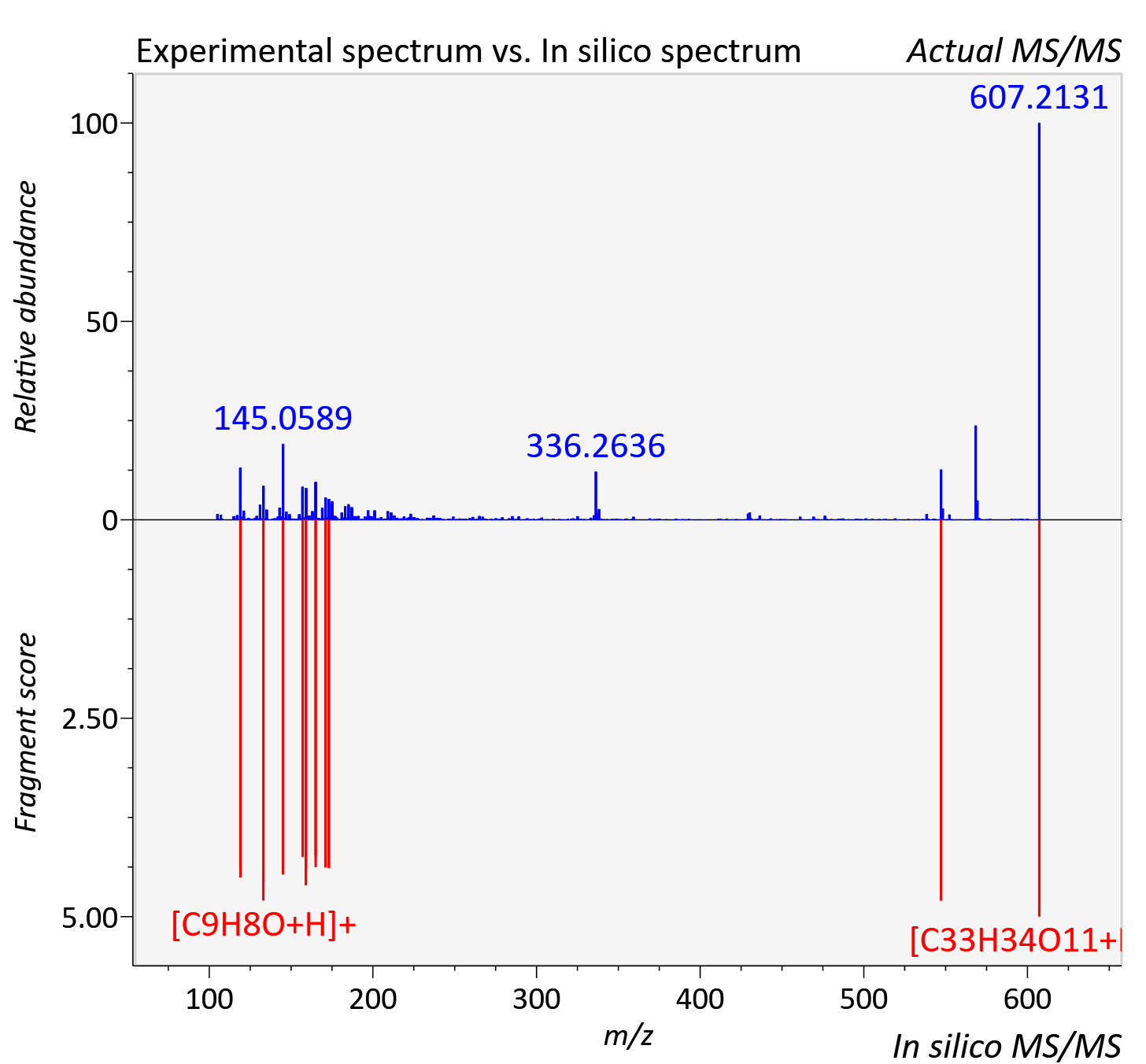
Experimental data
| Retention time | 14.22 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 607.213 | Theoretical mz | 607.217 |
| Error | 7.7 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.2384 |
Identifiers and class information
| Inchi key | JCDVQBGWJDNTFK-OUKQBFOZNA-N |
|---|---|
| Smiles | O=C(OC1C2=CC=3OCOC3C(OC)=C2C=4C(O)=C(OC)C(OC)=CC4C(OC(=O)C)C(O)(C)C1C)C=CC=5C=CC=CC5 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Tannins |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T25258 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08183 | ABCB1 |
| T23499 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P25101 | EDNRA |
| T23499 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P25101 | EDNRA |
| T23499 | DI0425 | Urinary system clinical symptom | [ICD-11: MF8Y] | P25101 | EDNRA |
| T92828 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P24530 | EDNRB |
| T92828 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P24530 | EDNRB |
| T86552 | DI0060 | Brain cancer | [ICD-11: 2A00] | P19438 | TNFRSF1A |
| T86552 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P19438 | TNFRSF1A |
