Compound details
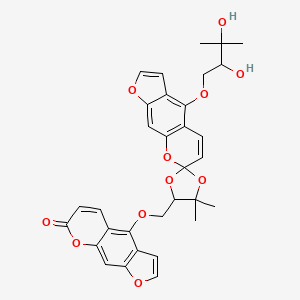
Dahuribirin E
| Compound ID | CDAMM00542 |
|---|---|
| Common name | Dahuribirin E | IUPAC name | 4-[[4'-(2,3-dihydroxy-3-methylbutoxy)-5,5-dimethylspiro[1,3-dioxolane-2,7'-furo[3,2-g]chromene]-4-yl]methoxy]furo[3,2-g]chromen-7-one |
| Molecular formula | C32H30O11 |
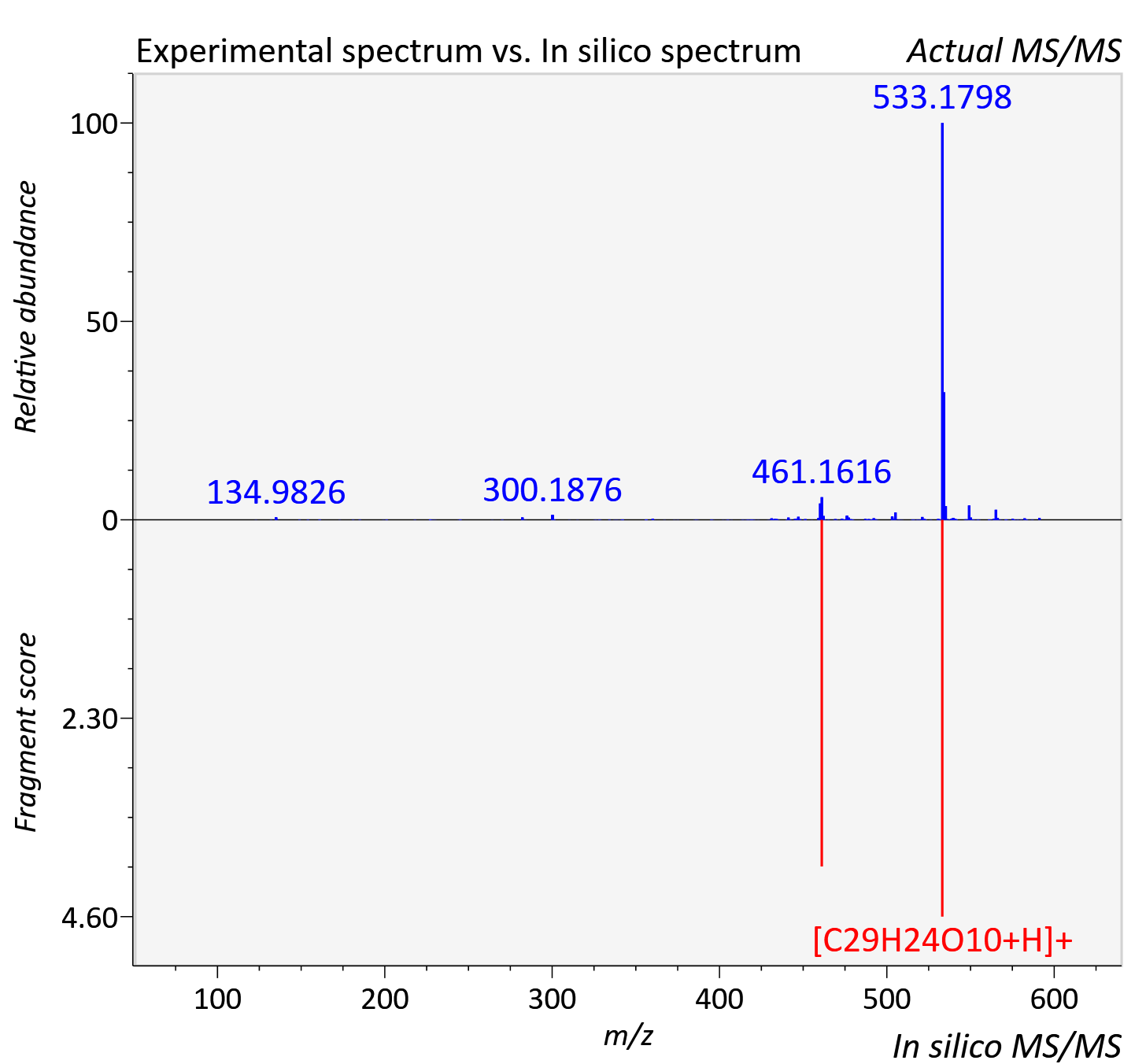
Experimental data
| Retention time | 13.29 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 591.185 | Theoretical mz | 591.186 |
| Error | 2.33 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.6064 |
Identifiers and class information
| Inchi key | JYGBUNBAQUBPQH-XCLNOZGTNA-N |
|---|---|
| Smiles | O=C1OC=2C=C3OC=CC3=C(OCC4OC5(OC=6C=C7OC=CC7=C(OCC(O)C(O)(C)C)C6C=C5)OC4(C)C)C2C=C1 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Coumarins and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P22460 | KCNA5 | Voltage-gated potassium channel subunit Kv1.5 | T17569 | SEA |
| P22001 | KCNA3 | Voltage-gated potassium channel subunit Kv1.3 | T76914 | SEA |
| Q92952 | KCNN1 | Small conductance calcium-activated potassium channel protein 1 | T11911 | SEA |
| Q9H2S1 | KCNN2 | Small conductance calcium-activated potassium channel protein 2 | T86271 | SEA |
| Q09470 | KCNA1 | Voltage-gated potassium channel Kv1.1 | T85272 | SEA |
| Q96RP8 | KCNA7 | Voltage-gated potassium channel Kv1.7 | T84737 | SEA |
| P22459 | KCNA4 | Voltage-gated A-type potassium channel | T15665 | SEA |
| P16389 | KCNA2 | Voltage-gated potassium channel Kv1.2 | T19492 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T17569 | DI0030 | Angina pectoris | [ICD-11: BA40] | P22460 | KCNA5 |
| T76914 | DI0198 | Idiopathic inflammatory myopathy | [ICD-11: 4A41] | P22001 | KCNA3 |
| T76914 | DI0351 | Psoriasis | [ICD-11: EA90] | P22001 | KCNA3 |
| T76914 | DI0352 | Psoriatic arthritis | [ICD-11: FA21] | P22001 | KCNA3 |
| T11911 | DI0285 | Myelopathy | [ICD-11: 8B42] | Q92952 | KCNN1 |
| T86271 | DI0285 | Myelopathy | [ICD-11: 8B42] | Q9H2S1 | KCNN2 |
