Compound details
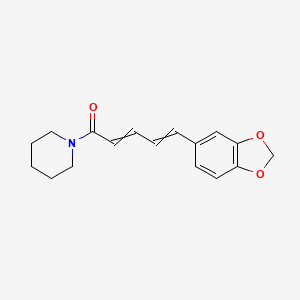
Piperine
| Compound ID | CDAMM00488 |
|---|---|
| Common name | Piperine | IUPAC name | 5-(1,3-benzodioxol-5-yl)-1-piperidin-1-ylpenta-2,4-dien-1-one |
| Molecular formula | C17H19NO3 |
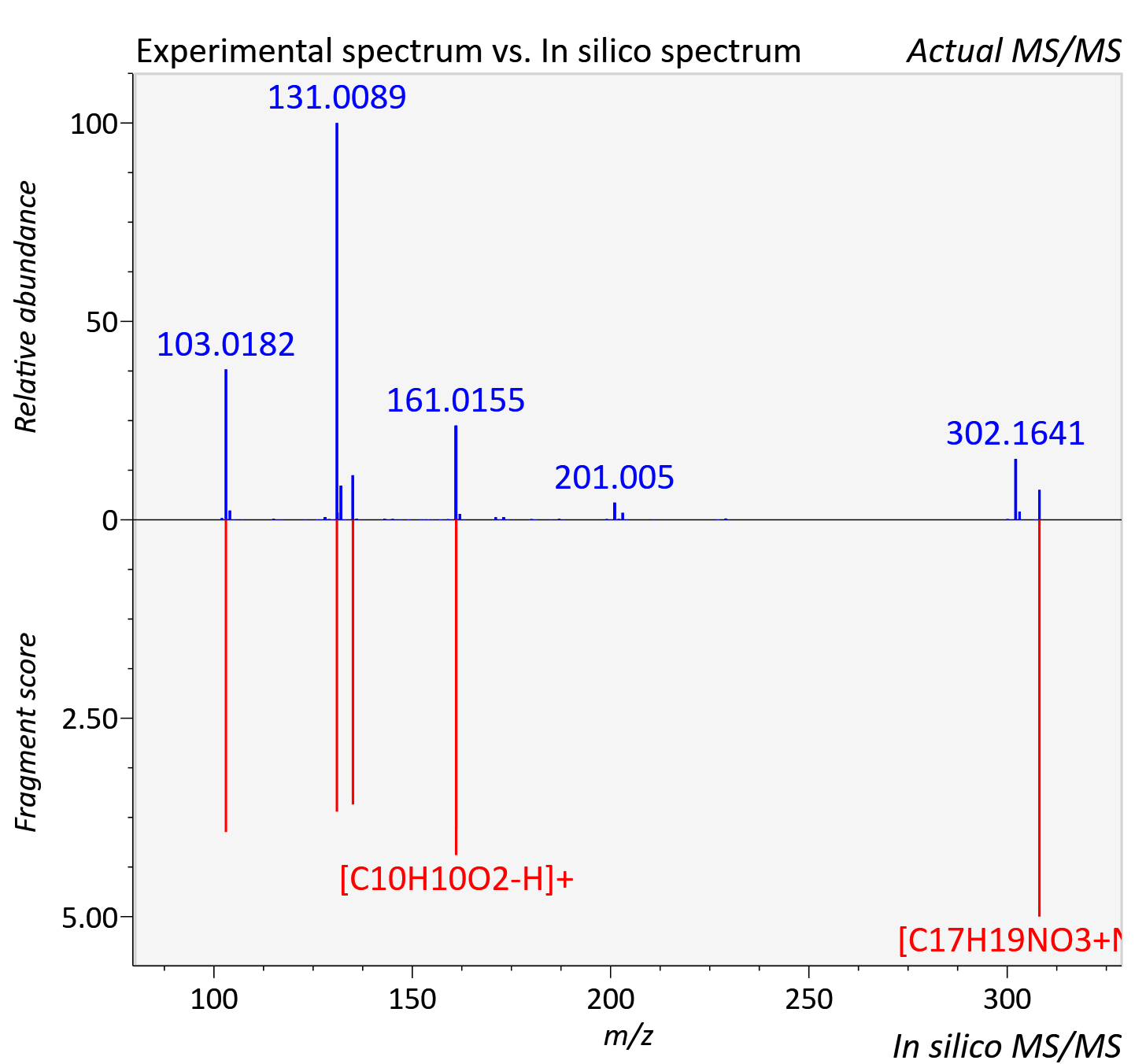
Experimental data
| Retention time | 8.06 |
|---|---|
| Adduct | [M+Na]+ |
| Actual mz | 308.125 | Theoretical mz | 308.125 |
| Error | 2.33 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.6408 |
Identifiers and class information
| Inchi key | MXXWOMGUGJBKIW-YPCIICBESA-N |
|---|---|
| Smiles | O=C(C=CC=CC1=CC=C2OCOC2=C1)N3CCCCC3 |
| Superclass | Alkaloids and derivatives |
| Class |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q99720 | SIGMAR1 | Sigma opioid receptor | T46360 | SwissTargetPrediction |
| Q16739 | UGCG | Ceramide glucosyltransferase | T14908 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SwissTargetPrediction and SEA |
| Q8NER1 | TRPV1 | Vanilloid receptor | T83193 | SEA |
| P08172 | CHRM2 | Muscarinic acetylcholine receptor M2 | T46185 | SEA |
| P20701 | ITGAL | Intercellular adhesion molecule (ICAM-1), Integrin alpha-L/beta-2 | T35640 | SEA |
| Q99685 | MGLL | Monoglyceride lipase | T18664 | SEA |
| P05362 | ICAM1 | Intercellular adhesion molecule-1 | T26203 | SEA |
| Q15078 | CDK5R1 | Cyclin-dependent kinase 5/CDK5 activator 1 | T09513 | SEA |
| P25101 | EDNRA | Endothelin receptor ET-A | T23499 | SEA |
| Q00535 | CDK5 | Cyclin-dependent kinase 5/CDK5 activator 1 | T20973 | SEA |
| P17936 | IGFBP3 | Insulin-like growth factor binding protein 3 | T33455 | SEA |
| O60344 | ECE2 | Endothelin-converting enzyme 2 | T16799 | SEA |
| Q92630 | DYRK2 | Dual-specificity tyrosine-phosphorylation regulated kinase 2 | T39486 | SEA |
| P19438 | TNFRSF1A | Tumor necrosis factor receptor R1 | T86552 | SEA |
| P32249 | EBI2 | EBV-induced G-protein coupled receptor 2 | T59577 | SEA |
| P51843 | NR0B1 | Orphan nuclear receptor DAX-1 | T23191 | SEA |
| Q7RTX1 | TAS1R1 | Taste receptor | T41263 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
| P05107 | ITGB2 | Integrin beta-2 | T10513 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T46360 | DI0105 | Cough | [ICD-11: MD12] | Q99720 | SIGMAR1 |
| T14908 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q16739 | UGCG |
| T14908 | DI0257 | Metabolic disorder | [ICD-11: 5C50-5D2Z] | Q16739 | UGCG |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T83193 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q8NER1 | TRPV1 |
| T46185 | DI0037 | Asthma | [ICD-11: CA23] | P08172 | CHRM2 |
| T46185 | DI0166 | Glaucoma | [ICD-11: 9C61] | P08172 | CHRM2 |
| T46185 | DI0278 | Muscle disorder | [ICD-11: FB32-FB3Z] | P08172 | CHRM2 |
| T46185 | DI0333 | Peptic ulcer | [ICD-11: DA61] | P08172 | CHRM2 |
| T46185 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08172 | CHRM2 |
| T46185 | DI0371 | Sebaceous gland disorder | [ICD-11: ED91] | P08172 | CHRM2 |
| T35640 | DI0351 | Psoriasis | [ICD-11: EA90] | P20701 | ITGAL |
| T35640 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P20701 | ITGAL |
| T18664 | DI0163 | General pain disorder | [ICD-11: 8E43] | Q99685 | MGLL |
| T18664 | DI0409 | Tic disorder | [ICD-11: 8A05] | Q99685 | MGLL |
| T26203 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P05362 | ICAM1 |
| T23499 | DI0070 | Cardiovascular disease | [ICD-11: BA00-BE2Z] | P25101 | EDNRA |
| T23499 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | P25101 | EDNRA |
| T23499 | DI0425 | Urinary system clinical symptom | [ICD-11: MF8Y] | P25101 | EDNRA |
| T20973 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | Q00535 | CDK5 |
| T33455 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P17936 | IGFBP3 |
| T86552 | DI0060 | Brain cancer | [ICD-11: 2A00] | P19438 | TNFRSF1A |
| T86552 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P19438 | TNFRSF1A |
| T41263 | DI0078 | Cholera | [ICD-11: 1A00] | Q7RTX1 | TAS1R1 |
| T10513 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P05107 | ITGB2 |
