Compound details
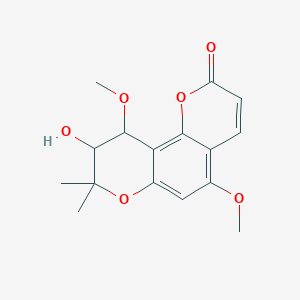
trans-O-Methylgrandmarin
| Compound ID | CDAMM00487 |
|---|---|
| Common name | trans-O-Methylgrandmarin | IUPAC name | 9-hydroxy-5,10-dimethoxy-8,8-dimethyl-9,10-dihydropyrano[2,3-f]chromen-2-one |
| Molecular formula | C16H18O6 |
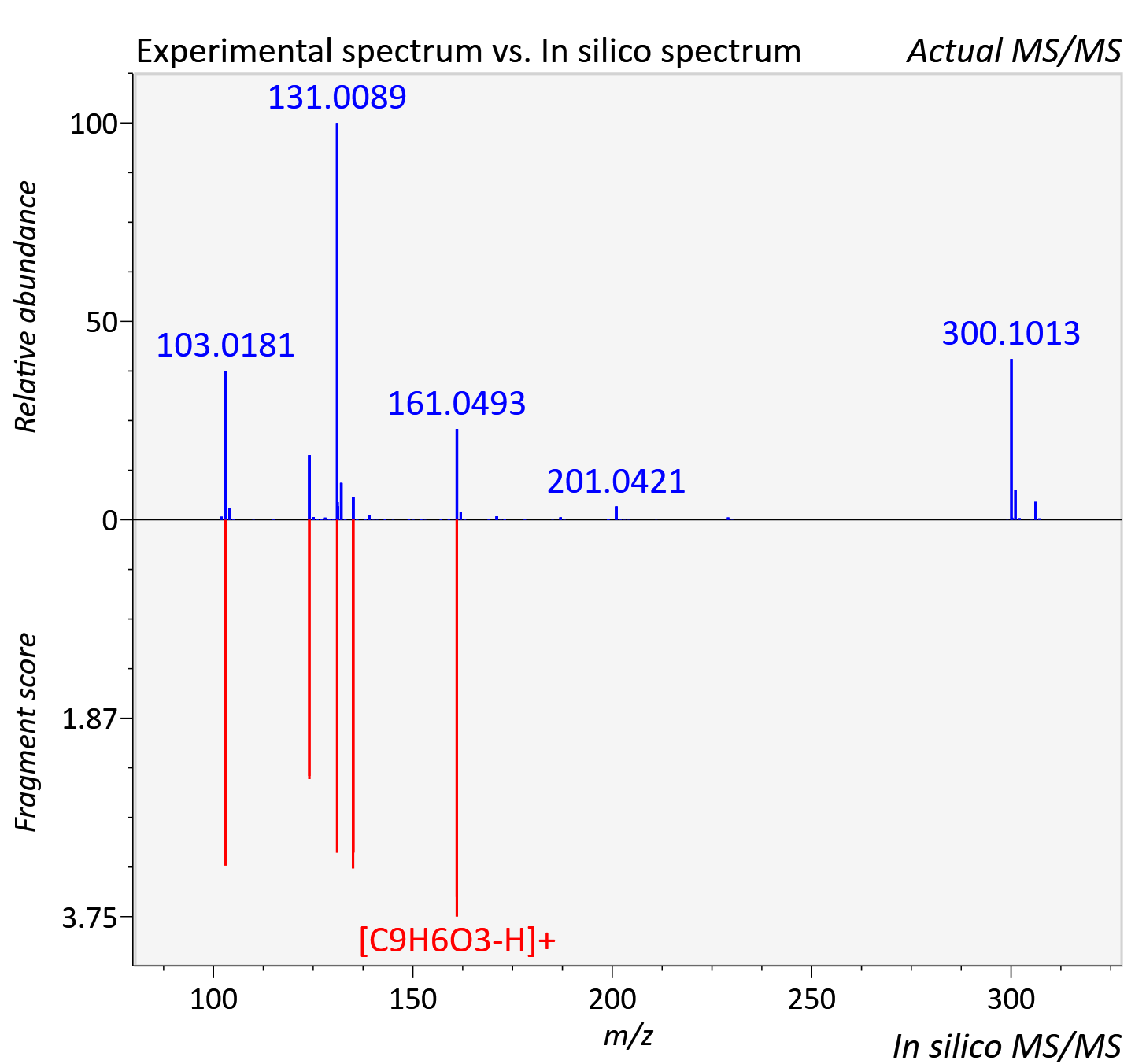
Experimental data
| Retention time | 7.63 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 307.114 | Theoretical mz | 307.117 |
| Error | 11.97 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.6384 |
Identifiers and class information
| Inchi key | NACAFQOYZHTCHZ-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1OC2=C(C=C1)C(OC)=CC=3OC(C)(C)C(O)C(OC)C32 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Coumarins and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T02001 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | Q08499 | PDE4D |
| T02777 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | O60706 | ABCC9 |
| T02777 | DI0030 | Angina pectoris | [ICD-11: BA40] | O60706 | ABCC9 |
| T02777 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | O60706 | ABCC9 |
