Compound details
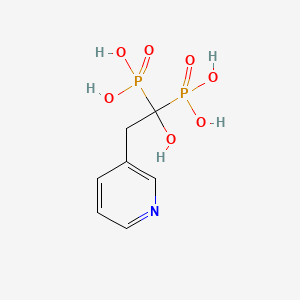
Risedronate
| Compound ID | CDAMM00482 |
|---|---|
| Common name | Risedronate | IUPAC name | (1-hydroxy-1-phosphono-2-pyridin-3-ylethyl)phosphonic acid |
| Molecular formula | C7H11NO7P2 |
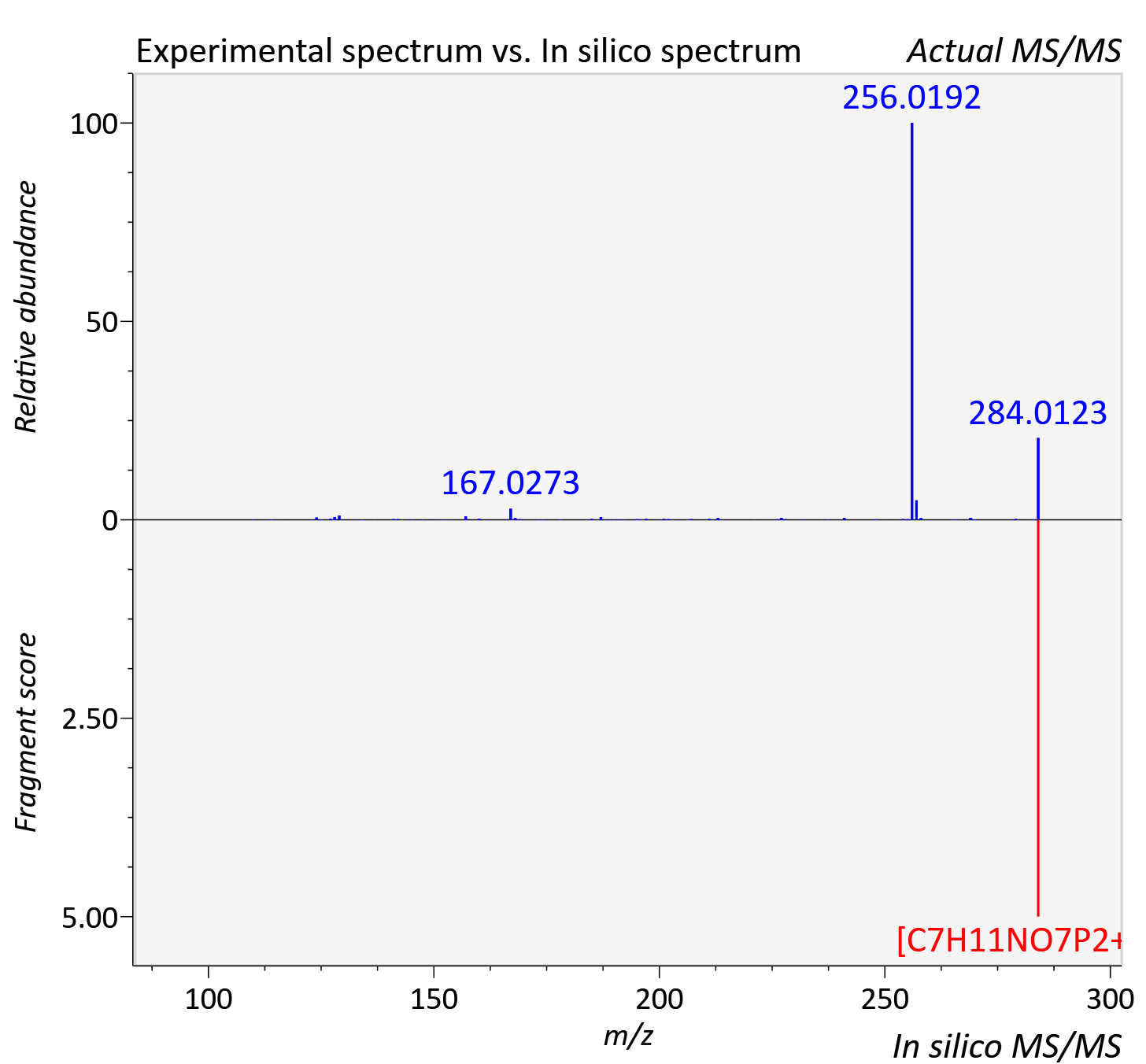
Experimental data
| Retention time | 8.81 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 284.012 | Theoretical mz | 284.008 |
| Error | 11.7 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.5377 |
Identifiers and class information
| Inchi key | IIDJRNMFWXDHID-UHFFFAOYSA-N |
|---|---|
| Smiles | O=P(O)(O)C(O)(CC=1C=NC=CC1)P(=O)(O)O |
| Superclass | Organic acids and derivatives |
| Class | Organic phosphonic acids and derivatives |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P43681 | CHRNA4 | Neuronal acetylcholine receptor; alpha4/beta2 | T70967 | SEA |
| P36544 | CHRNA7 | Neuronal acetylcholine receptor protein alpha-7 subunit | T34429 | SEA |
| P15144 | ANPEP | Aminopeptidase N | T67272 | SEA |
| P11509 | CYP2A6 | Cytochrome P450 2A6 | T06455 | SEA |
| Q14416 | GRM2 | Metabotropic glutamate receptor 2 | T62820 | SEA |
| P15538 | CYP11B1 | Cytochrome P450 11B1 | T84621 | SEA |
| P25105 | PTAFR | Platelet activating factor receptor | T87023 | SEA |
| Q16581 | C3AR1 | C3a anaphylatoxin chemotactic receptor | T30219 | SEA |
| P43490 | NAMPT | Nicotinamide phosphoribosyltransferase | T54582 | SEA |
| P17787 | CHRNB2 | Nicotinic acetylcholine receptor alpha2/beta2 | T82543 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SwissTargetPrediction and SEA |
| P24557 | TBXAS1 | Thromboxane-A synthase | T78356 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| Q9UG96 | ICH2 | Caspase-4 | T53469 | SEA |
| Q6DJV7 | ICH3 | Caspase-5 | T89565 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T70967 | DI0029 | Aneurysm/dissection | [ICD-11: BD50] | P43681 | CHRNA4 |
| T70967 | DI0191 | Hypertensive crisis | [ICD-11: BA03] | P43681 | CHRNA4 |
| T70967 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | P43681 | CHRNA4 |
| T34429 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P36544 | CHRNA7 |
| T34429 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P36544 | CHRNA7 |
| T67272 | DI0351 | Psoriasis | [ICD-11: EA90] | P15144 | ANPEP |
| T06455 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P11509 | CYP2A6 |
| T06455 | DI0351 | Psoriasis | [ICD-11: EA90] | P11509 | CYP2A6 |
| T62820 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q14416 | GRM2 |
| T62820 | DI0051 | Bipolar disorder | [ICD-11: 6A60] | Q14416 | GRM2 |
| T62820 | DI0116 | Dengue fever | [ICD-11: 1D2Z] | Q14416 | GRM2 |
| T62820 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | Q14416 | GRM2 |
| T62820 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | Q14416 | GRM2 |
| T62820 | DI0256 | Mental/behavioural/neurodevelopmental disorder | [ICD-11: 6E20-6E8Z] | Q14416 | GRM2 |
| T62820 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | Q14416 | GRM2 |
| T62820 | DI0354 | Psychotic disorder | [ICD-11: 6A20-6A25] | Q14416 | GRM2 |
| T62820 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q14416 | GRM2 |
| T84621 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P15538 | CYP11B1 |
| T84621 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P15538 | CYP11B1 |
| T87023 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P25105 | PTAFR |
| T54582 | DI0231 | Leukaemia | [ICD-11: 2A60-2B33] | P43490 | NAMPT |
| T54582 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P43490 | NAMPT |
| T54582 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P43490 | NAMPT |
| T82543 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P17787 | CHRNB2 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T78356 | DI0030 | Angina pectoris | [ICD-11: BA40] | P24557 | TBXAS1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
