Compound details
m-Cresol
| Compound ID | CDAMM00453 |
|---|---|
| Common name | m-Cresol | IUPAC name | 3-methylphenol |
| Molecular formula | C7H8O |
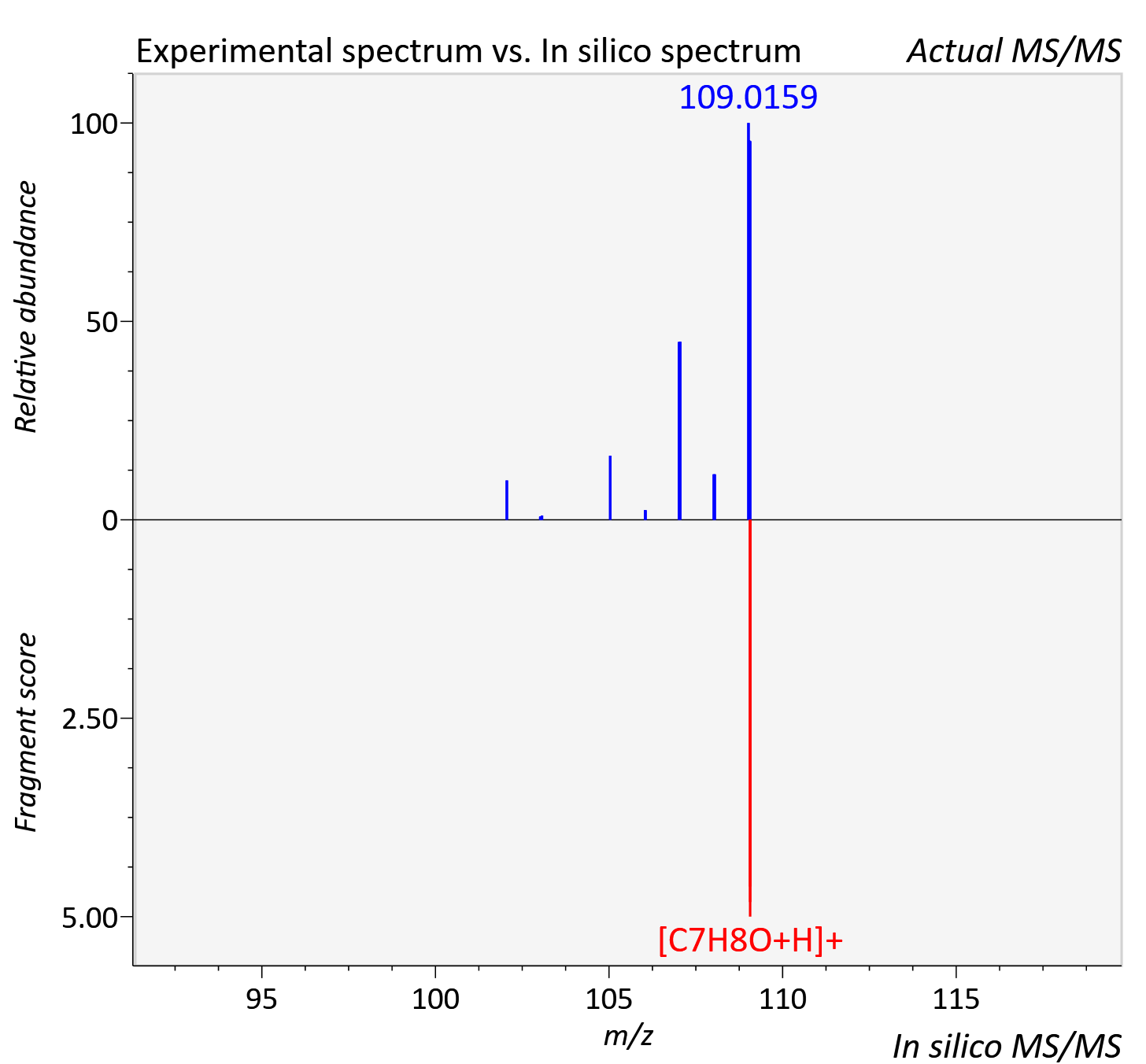
Experimental data
| Retention time | 13.51 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 109.065 | Theoretical mz | 109.065 |
| Error | 3.45 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.1683 |
Identifiers and class information
| Inchi key | RLSSMJSEOOYNOY-UHFFFAOYSA-N |
|---|---|
| Smiles | OC1=CC=CC(=C1)C |
| Superclass | Benzenoids |
| Class | Phenols |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P22303 | ACHE | Acetylcholinesterase | T30082 | SwissTargetPrediction and SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P27338 | MAOB | Monoamine oxidase B | T83011 | SEA |
| P32297 | CHRNA3 | Neuronal acetylcholine receptor; alpha3/beta4 | T74166 | SEA |
| P14920 | DAO | D-amino-acid oxidase | T33124 | SEA |
| Q92731 | ESR2 | Estrogen receptor beta | T80896 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SEA |
| P03372 | ESR1 | Estrogen receptor alpha | T02506 | SEA |
| P09467 | FBP1 | Fructose-1,6-bisphosphatase | T83391 | SEA |
| P16050 | ALOX15 | Arachidonate 15-lipoxygenase | T16042 | SEA |
| Q99523 | SORT1 | Neurotensin receptor 3 | T63233 | SEA |
| O14649 | KCNK3 | Potassium channel subfamily K member 3 | T15795 | SEA |
| Q9NPC2 | KCNK9 | Potassium channel subfamily K member 9 | T57613 | SEA |
| O95747 | OXSR1 | Serine/threonine-protein kinase OSR1 | T08198 | SEA |
| Q8WZA2 | RAPGEF4 | Rap guanine nucleotide exchange factor 4 | T74607 | SEA |
| P30926 | CHRNB4 | Neuronal acetylcholine receptor; alpha2/beta4 | T73724 | SEA |
| P35270 | SPR | Substance-P receptor | T47094 | SEA |
| Q9C000 | NLRP1 | NLR pyrin domain containing 1 | T86910 | SEA |
| P02708 | CHRNA1 | Neuronal acetylcholine receptor alpha-1 | T04689 | SEA |
| Q9BX84 | TRPM6 | Long transient receptor potential channel 6 | T61029 | SEA |
| Q8IWQ3 | BRSK2 | BR serine/threonine kinase 2 | T10375 | SEA |
| P57078 | RIPK4 | PKC-delta-interacting protein kinase | T93594 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T30082 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P22303 | ACHE |
| T30082 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22303 | ACHE |
| T30082 | DI0282 | Myasthenia gravis | [ICD-11: 8C6Y] | P22303 | ACHE |
| T30082 | DI0313 | Oesophageal/gastroduodenal disorder | [ICD-11: DD90] | P22303 | ACHE |
| T30082 | DI0332 | Pediculosis | [ICD-11: 1G00] | P22303 | ACHE |
| T30082 | DI0421 | Unspecific substance harmful effect | [ICD-11: NE6Z] | P22303 | ACHE |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T83011 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | P27338 | MAOB |
| T83011 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P27338 | MAOB |
| T83011 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P27338 | MAOB |
| T83011 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P27338 | MAOB |
| T83011 | DI0264 | Migraine | [ICD-11: 8A80] | P27338 | MAOB |
| T83011 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P27338 | MAOB |
| T74166 | DI0105 | Cough | [ICD-11: MD12] | P32297 | CHRNA3 |
| T33124 | DI0038 | Ataxic disorder | [ICD-11: 8A03] | P14920 | DAO |
| T33124 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P14920 | DAO |
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T83391 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P09467 | FBP1 |
| T63233 | DI0153 | Frontotemporal dementia | [ICD-11: 6D83] | Q99523 | SORT1 |
| T15795 | DI0309 | Obstructive sleep apnoea | [ICD-11: 7A41] | O14649 | KCNK3 |
| T73724 | DI0301 | Nicotine use disorder | [ICD-11: 6C4A] | P30926 | CHRNB4 |
| T47094 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P35270 | SPR |
| T47094 | DI0293 | Nausea/vomiting | [ICD-11: MD90] | P35270 | SPR |
| T04689 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P02708 | CHRNA1 |
