Compound details
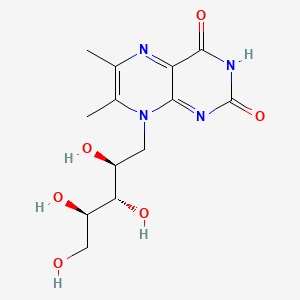
6,7-Dimethyl-8-(1-D-ribityl)lumazine
| Compound ID | CDAMM00412 |
|---|---|
| Common name | 6,7-Dimethyl-8-(1-D-ribityl)lumazine | IUPAC name | 6,7-dimethyl-8-[(2S,3S,4R)-2,3,4,5-tetrahydroxypentyl]pteridine-2,4-dione |
| Molecular formula | C13H18N4O6 |
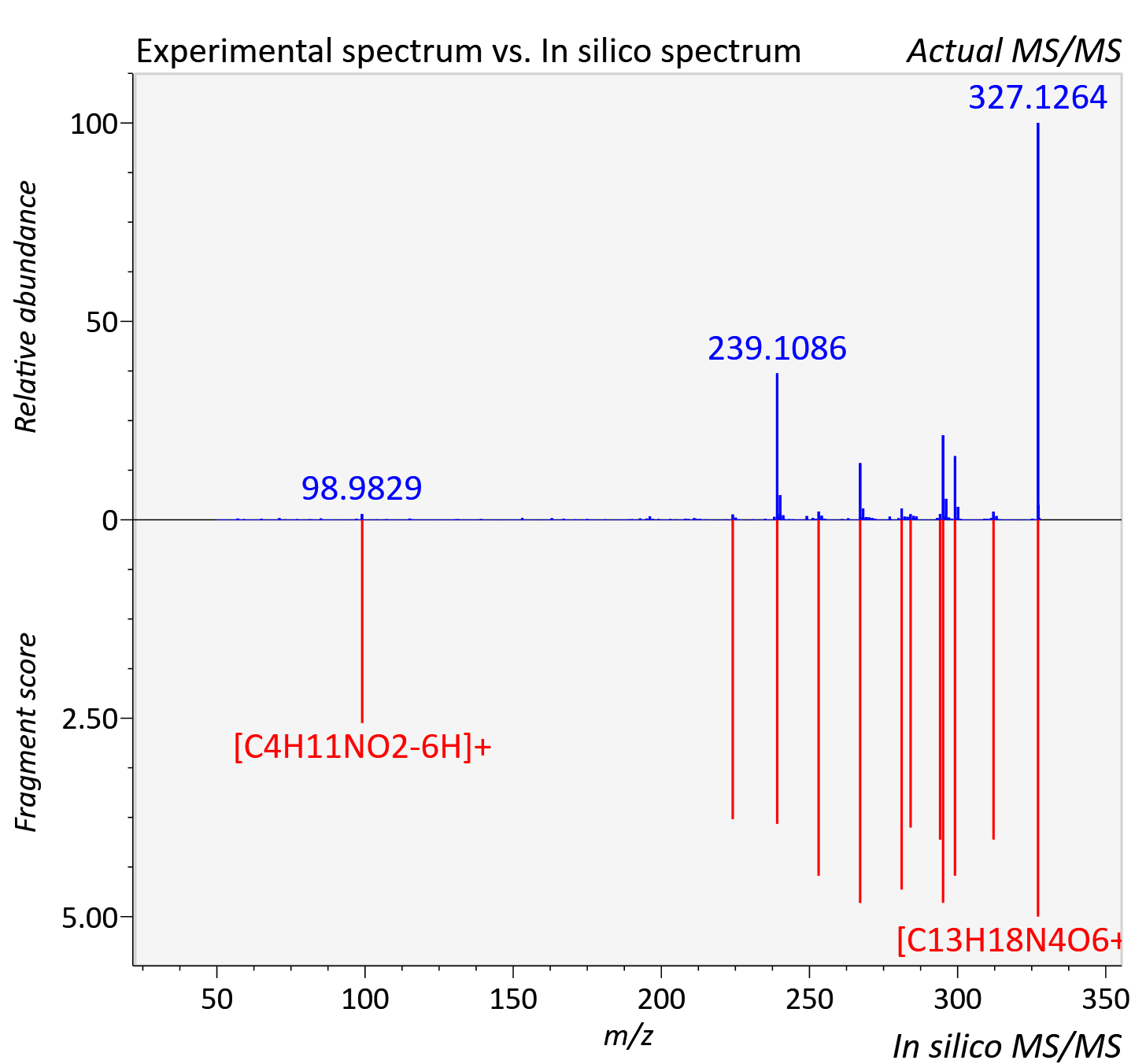
Experimental data
| Retention time | 61.82 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 327.126 | Theoretical mz | 327.13 |
| Error | 13.17 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.53 |
Identifiers and class information
| Inchi key | SXDXRJZUAJBNFL-XKSSXDPKSA-N |
|---|---|
| Smiles | CC1=C(N(C2=NC(=O)NC(=O)C2=N1)C[C@@H]([C@@H]([C@@H](CO)O)O)O)C |
| Superclass | Organoheterocyclic compounds |
| Class | Pteridines and derivatives |
