Compound details
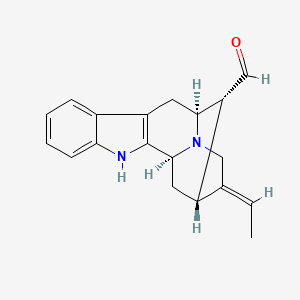
Vellosimine
| Compound ID | CDAMM00409 |
|---|---|
| Common name | Vellosimine | IUPAC name | (1S,12S,13R,14R,15E)-15-ethylidene-3,17-diazapentacyclo[12.3.1.02,10.04,9.012,17]octadeca-2(10),4,6,8-tetraene-13-carbaldehyde |
| Molecular formula | C19H20N2O |
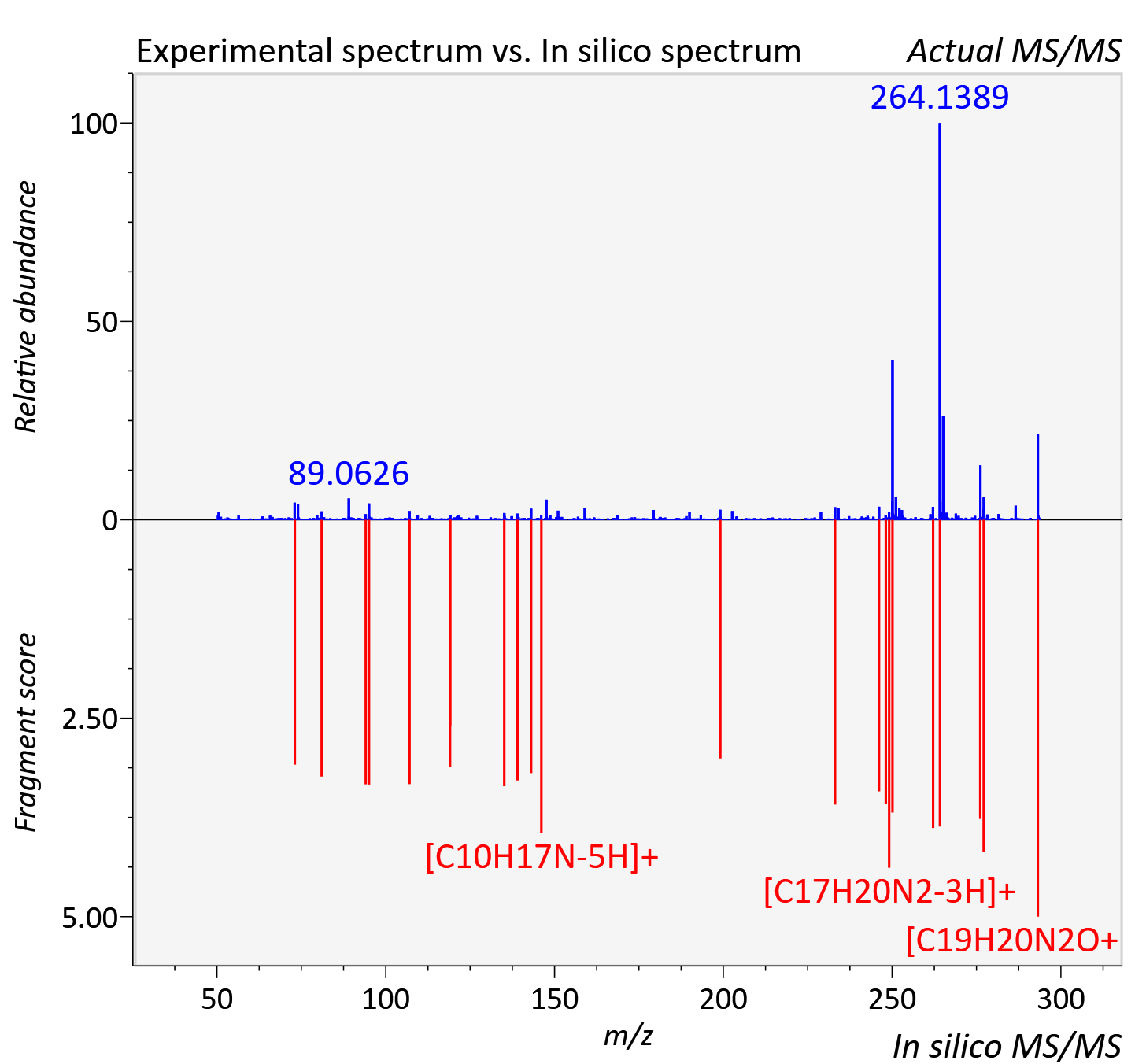
Experimental data
| Retention time | 44.817 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 293.168 | Theoretical mz | 293.165 |
| Error | 9.41 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.26 |
Identifiers and class information
| Inchi key | MHASSCPGKAMILD-YALZJJMTNA-N |
|---|---|
| Smiles | C/C=C\1/CN2[C@H]3C[C@@H]1[C@H]([C@@H]2CC4=C3NC5=CC=CC=C45)C=O |
| Superclass | Alkaloids and derivatives |
| Class | Macroline alkaloids |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T07663 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | O76074 | PDE5A |
| T07663 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | O76074 | PDE5A |
| T07663 | DI0378 | Sexual dysfunction | [ICD-11: HA00-HA01] | O76074 | PDE5A |
| T07663 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | O76074 | PDE5A |
| T60514 | DI0035 | Arterial occlusive disease | [ICD-11: BD40] | Q9HCN6 | GP6 |
| T60514 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q9HCN6 | GP6 |
