Compound details
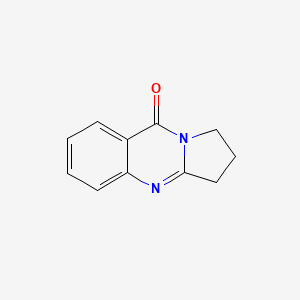
Deoxyvasicinone
| Compound ID | CDAMM00405 |
|---|---|
| Common name | Deoxyvasicinone | IUPAC name | 2,3-dihydro-1H-pyrrolo[2,1-b]quinazolin-9-one |
| Molecular formula | C11H10N2O |
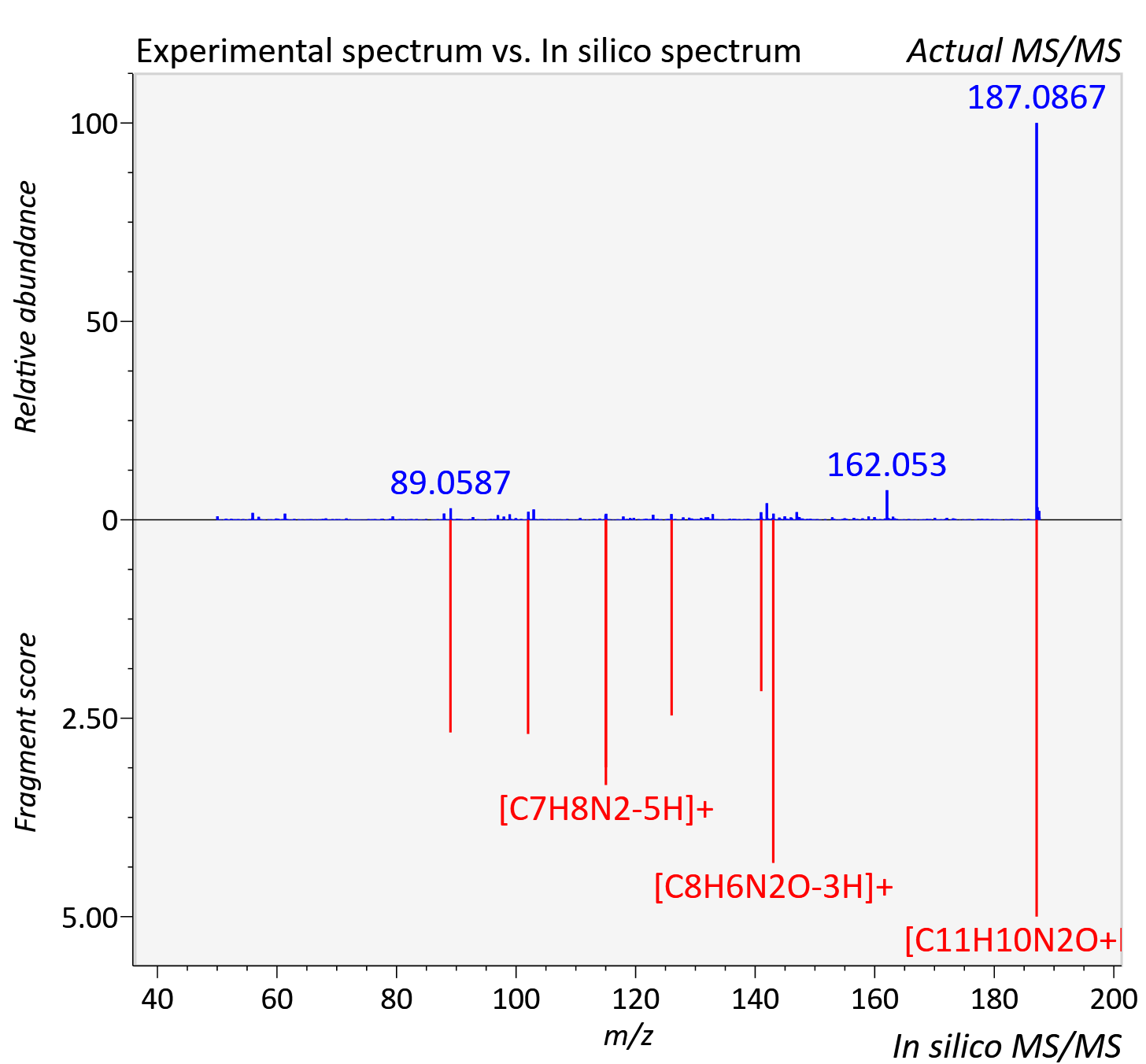
Experimental data
| Retention time | 20.842 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 187.088 | Theoretical mz | 187.087 |
| Error | 6.47 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.66 |
Identifiers and class information
| Inchi key | VARHXCYGZKSOOO-UHFFFAOYSA-N |
|---|---|
| Smiles | C1CC2=NC3=CC=CC=C3C(=O)N2C1 |
| Superclass | Organoheterocyclic compounds |
| Class | Diazanaphthalenes |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q9Y5N1 | HRH3 | Histamine H3 receptor | T64765 | SEA |
| P22303 | ACHE | Acetylcholinesterase | T30082 | SEA |
| P34969 | HTR7 | Serotonin 7 (5-HT7) receptor | T79062 | SEA |
| P05164 | MPO | Myeloperoxidase | T23471 | SEA |
| P33527 | ABCC1 | Multidrug resistance-associated protein 1 | T11288 | SEA |
| O95271 | TNKS | Tankyrase-1 | T83059 | SEA |
| P06276 | BCHE | Butyrylcholinesterase | T99799 | SEA |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| P08236 | GUSB | Beta-glucuronidase | T96413 | SEA |
| P42224 | STAT1 | Signal transducer and activator of transcription 1-alpha/beta | T64205 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| P53582 | METAP1 | Methionine aminopeptidase 1 | T42337 | SEA |
| Q13564 | NAE1 | NEDD8-activating enzyme E1 regulatory subunit | T01447 | SEA |
| P51532 | SMARCA4 | Transcription activator BRG1 | T69200 | SEA |
| Q86U86 | PBRM1 | Protein polybromo-1 | T00973 | SEA |
| O95602 | POLR1A | DNA-directed RNA polymerase I subunit RPA1 | T89530 | SEA |
| Q06609 | RAD51 | DNA repair protein RAD51 homolog 1 | T63083 | SEA |
| Q6ZQW0 | INDOL1 | Indoleamine 2,3-dioxygenase 2 | T62266 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T64765 | DI0392 | Somnolence | [ICD-11: MG42] | Q9Y5N1 | HRH3 |
| T30082 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P22303 | ACHE |
| T30082 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22303 | ACHE |
| T30082 | DI0282 | Myasthenia gravis | [ICD-11: 8C6Y] | P22303 | ACHE |
| T30082 | DI0313 | Oesophageal/gastroduodenal disorder | [ICD-11: DD90] | P22303 | ACHE |
| T30082 | DI0332 | Pediculosis | [ICD-11: 1G00] | P22303 | ACHE |
| T30082 | DI0421 | Unspecific substance harmful effect | [ICD-11: NE6Z] | P22303 | ACHE |
| T79062 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | P34969 | HTR7 |
| T79062 | DI0040 | Attention deficit hyperactivity disorder | [ICD-11: 6A05] | P34969 | HTR7 |
| T79062 | DI0051 | Bipolar disorder | [ICD-11: 6A60] | P34969 | HTR7 |
| T79062 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P34969 | HTR7 |
| T79062 | DI0265 | Mild neurocognitive disorder | [ICD-11: 6D71] | P34969 | HTR7 |
| T79062 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P34969 | HTR7 |
| T79062 | DI0370 | Schizophrenia | [ICD-11: 6A20] | P34969 | HTR7 |
| T23471 | DI0229 | Left ventricular failure | [ICD-11: BD11] | P05164 | MPO |
| T23471 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P05164 | MPO |
| T11288 | DI0167 | Gout | [ICD-11: FA25] | P33527 | ABCC1 |
| T83059 | DI0060 | Brain cancer | [ICD-11: 2A00] | O95271 | TNKS |
| T83059 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | O95271 | TNKS |
| T83059 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | O95271 | TNKS |
| T83059 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O95271 | TNKS |
| T99799 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P06276 | BCHE |
| T99799 | DI0411 | Tonus and reflex abnormality | [ICD-11: MB47] | P06276 | BCHE |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T96413 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | P08236 | GUSB |
| T64205 | DI0383 | Skin and skin-structure infection | [ICD-11: 1F28-1G0Z] | P42224 | STAT1 |
| T01447 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | Q13564 | NAE1 |
| T63083 | DI0048 | B-cell lymphoma | [ICD-11: 2A86] | Q06609 | RAD51 |
| T63083 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q06609 | RAD51 |
