Compound details
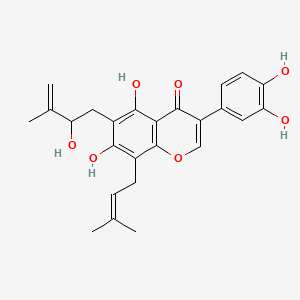
Millewanin G
| Compound ID | CDAMM00386 |
|---|---|
| Common name | Millewanin G | IUPAC name | 3-(3,4-dihydroxyphenyl)-5,7-dihydroxy-6-(2-hydroxy-3-methylbut-3-enyl)-8-(3-methylbut-2-enyl)chromen-4-one |
| Molecular formula | C25H26O7 |
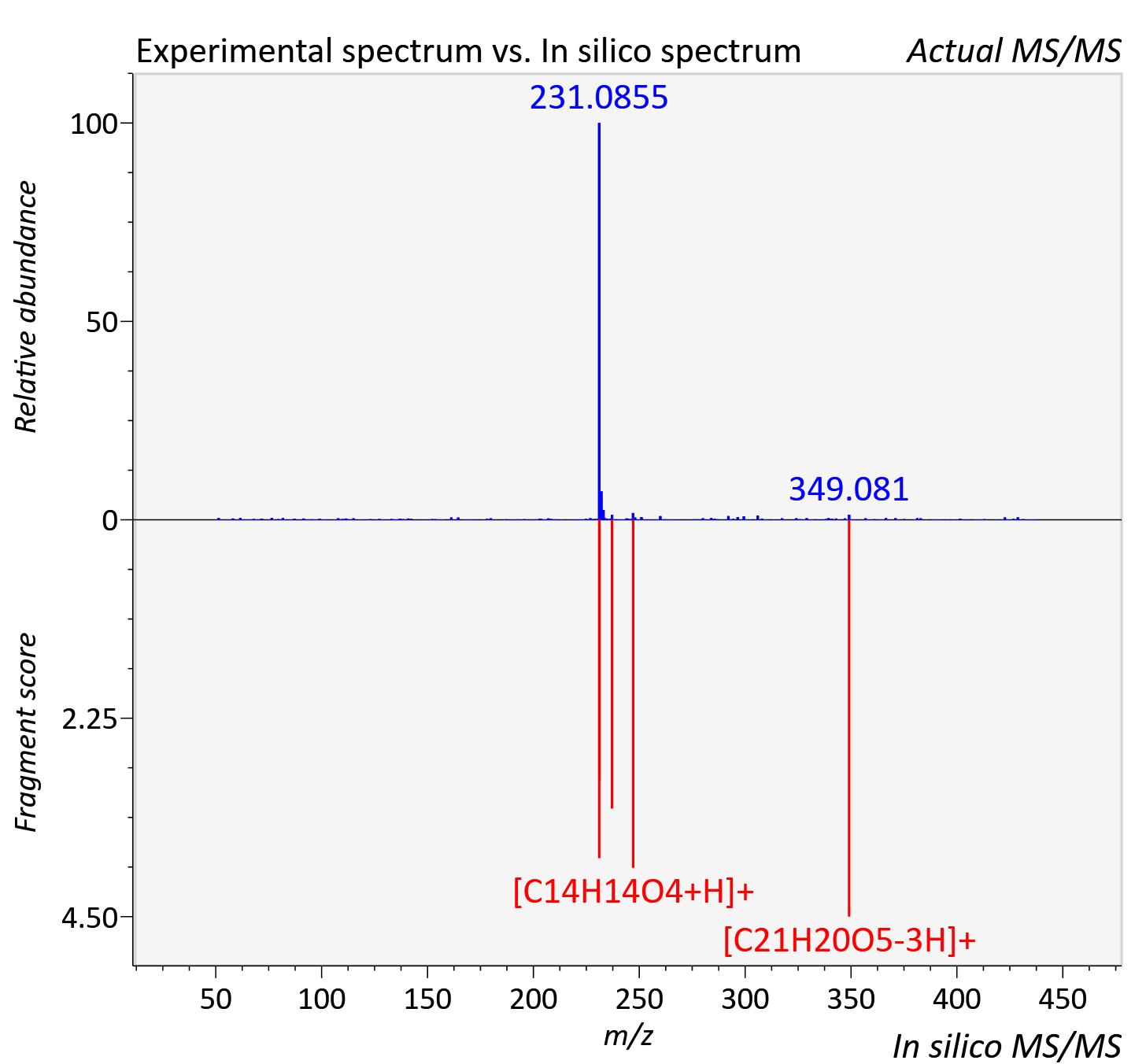
Experimental data
| Retention time | 9.665 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 439.178 | Theoretical mz | 439.175 |
| Error | 6.08 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.89 |
Identifiers and class information
| Inchi key | FSQKKJIBNQATIX-UHFFFAOYNA-N |
|---|---|
| Smiles | O=C1C(=COC2=C1C(O)=C(C(O)=C2CC=C(C)C)CC(O)C(=C)C)C=3C=CC(O)=C(O)C3 |
| Superclass | Phenylpropanoids and polyketides |
| Class | Isoflavonoids |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P00734 | F2 | Thrombin | T94033 | SwissTargetPrediction |
| P19838 | NFKB1 | Nuclear factor NF-kappa-B p105 subunit | T83145 | SEA |
| P18031 | PTPN1 | Protein-tyrosine phosphatase 1B | T16347 | SwissTargetPrediction and SEA |
| P18054 | ALOX12 | Arachidonate 12-lipoxygenase | T35527 | SEA |
| P00742 | F10 | Thrombin and coagulation factor X | T84631 | SwissTargetPrediction |
| P05771 | PRKCB | Protein kinase C beta | T40276 | SwissTargetPrediction |
| P06870 | KLK1 | Kallikrein 1 | T40000 | SwissTargetPrediction |
| P20151 | KLK2 | Kallikrein 2 | T01908 | SwissTargetPrediction |
| P08709 | F7 | Coagulation factor VII | T43332 | SwissTargetPrediction |
| P05129 | PRKCG | Protein kinase C gamma | T47107 | SwissTargetPrediction |
| O60911 | CTSV | Cathepsin (V and K) | T93653 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SEA |
| Q13332 | PTPRS | Receptor-type tyrosine-protein phosphatase S | T10147 | SEA |
| Q96HK3 | CALM | Calmodulin | T39610 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T94033 | DI0052 | Bleeding disorder | [ICD-11: GA20-GA21] | P00734 | F2 |
| T94033 | DI0091 | Coagulation defect | [ICD-11: 3B10] | P00734 | F2 |
| T94033 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P00734 | F2 |
| T94033 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P00734 | F2 |
| T94033 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P00734 | F2 |
| T94033 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P00734 | F2 |
| T94033 | DI0403 | Thrombocytopenia | [ICD-11: 3B64] | P00734 | F2 |
| T94033 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P00734 | F2 |
| T83145 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P19838 | NFKB1 |
| T83145 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P19838 | NFKB1 |
| T16347 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | P18031 | PTPN1 |
| T16347 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P18031 | PTPN1 |
| T16347 | DI0308 | Obesity | [ICD-11: 5B80-5B81] | P18031 | PTPN1 |
| T16347 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | P18031 | PTPN1 |
| T84631 | DI0053 | Blood-forming organ disease | [ICD-11: 3C0Z] | P00742 | F10 |
| T84631 | DI0080 | Christmas disease | [ICD-11: 3B11] | P00742 | F10 |
| T84631 | DI0091 | Coagulation defect | [ICD-11: 3B10] | P00742 | F10 |
| T84631 | DI0103 | Coronary thrombosis | [ICD-11: BA43] | P00742 | F10 |
| T84631 | DI0114 | Deep vein thrombosis | [ICD-11: BD71] | P00742 | F10 |
| T84631 | DI0397 | Supraventricular tachyarrhythmia | [ICD-11: BC81] | P00742 | F10 |
| T84631 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | P00742 | F10 |
| T84631 | DI0433 | Venous thromboembolism | [ICD-11: BD72] | P00742 | F10 |
| T40276 | DI0123 | Diffuse large B-cell lymphoma | [ICD-11: 2A81] | P05771 | PRKCB |
| T40276 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P05771 | PRKCB |
| T40276 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P05771 | PRKCB |
| T01908 | DI0213 | Innate/adaptive immunodeficiency | [ICD-11: 4A00] | P20151 | KLK2 |
| T43332 | DI0091 | Coagulation defect | [ICD-11: 3B10] | P08709 | F7 |
| T47107 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P05129 | PRKCG |
| T47107 | DI0248 | Mastocytosis | [ICD-11: 2A21] | P05129 | PRKCG |
| T93653 | DI0055 | Bone cancer | [ICD-11: 2B5Z] | O60911 | CTSV |
| T93653 | DI0087 | Chronic pain | [ICD-11: MG30] | O60911 | CTSV |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T10147 | DI0057 | Bone paget disease | [ICD-11: FB85] | Q13332 | PTPRS |
| T39610 | DI0068 | Cardiac arrhythmia | [ICD-11: BC9Z] | Q96HK3 | CALM |
| T39610 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | Q96HK3 | CALM |
| T39610 | DI0370 | Schizophrenia | [ICD-11: 6A20] | Q96HK3 | CALM |
