Compound details
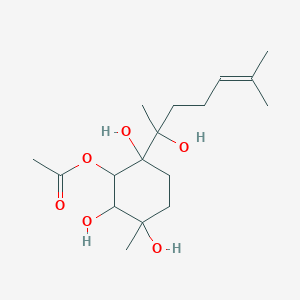
(1R,2R,3R ,6R,7R)-1,2,3,6,7-pentahydroxy-1-acetoxy-bisabol-10(11)-ene
| Compound ID | CDAMM00382 |
|---|---|
| Common name | (1R,2R,3R ,6R,7R)-1,2,3,6,7-pentahydroxy-1-acetoxy-bisabol-10(11)-ene | IUPAC name | [2,5,6-trihydroxy-2-(2-hydroxy-6-methylhept-5-en-2-yl)-5-methylcyclohexyl] acetate |
| Molecular formula | C17H30O6 |
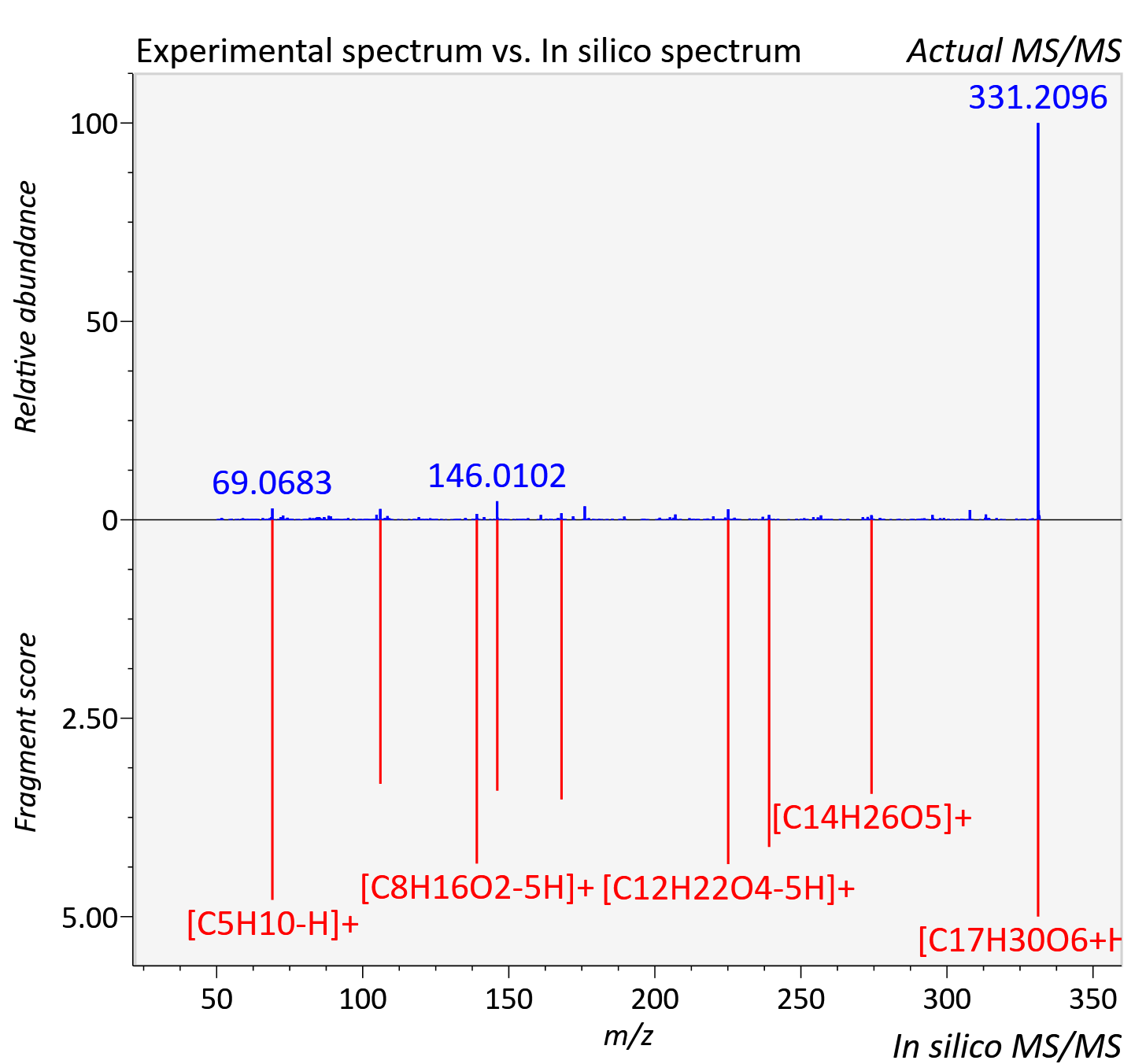
Experimental data
| Retention time | 51.24 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 331.21 | Theoretical mz | 331.201 |
| Error | 0 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.92 |
Identifiers and class information
| Inchi key | KNFIGEXUOBWKSE-WRQOLXDDSA-N |
|---|---|
| Smiles | O=C(OC1C(O)C(O)(C)CCC1(O)C(O)(C)CCC=C(C)C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Plant source
External sources
| HMDB ID | |
|---|---|
| KNApSAcK ID | |
| FooDB ID | |
| DrugBank ID | |
| ChEBI ID | |
| Pubchem Compound ID | 85258001 |
| PlantCyc ID | |
| UNPD ID | UNPD124596 |
| Coconut ID | |
| LipidMAPS ID | |
| NANPDB ID | NANPDB_6040 |
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T63609 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | Q08828 | ADCY1 |
| T96723 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | Q15393 | SF3B3 |
| T96723 | DI0085 | Chronic myelomonocytic leukaemia | [ICD-11: 2A40] | Q15393 | SF3B3 |
| T96723 | DI0284 | Myelodysplastic syndrome | [ICD-11: 2A37] | Q15393 | SF3B3 |
