Compound details
p-Cymene
| Compound ID | CDAMM00368 |
|---|---|
| Common name | p-Cymene | IUPAC name | 1-methyl-4-propan-2-ylbenzene |
| Molecular formula | C10H14 |
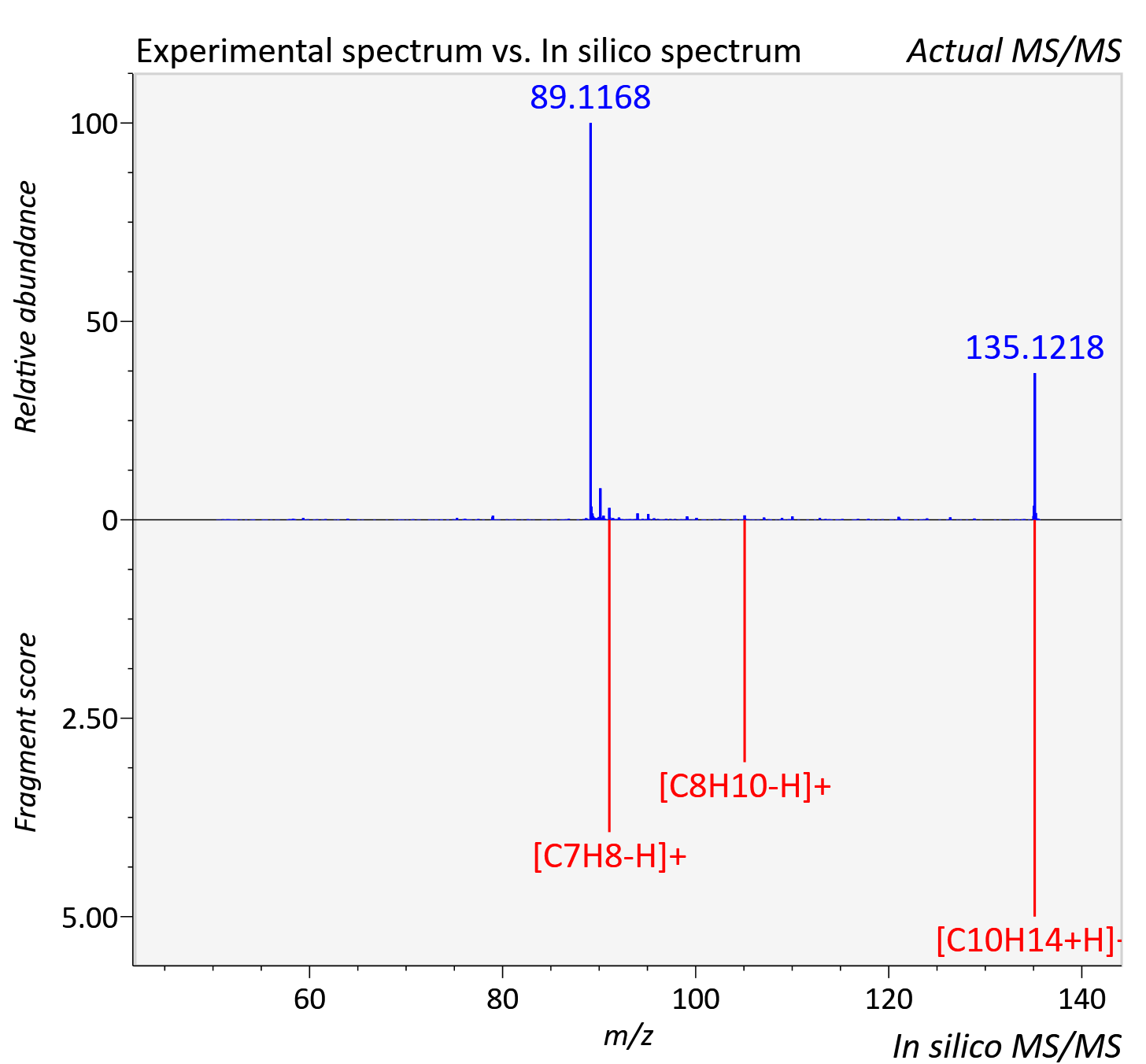
Experimental data
| Retention time | 12.992 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 135.121 | Theoretical mz | 135.117 |
| Error | 30.15 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.69 |
Identifiers and class information
| Inchi key | HFPZCAJZSCWRBC-UHFFFAOYSA-N |
|---|---|
| Smiles | C=1C=C(C=CC1C)C(C)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Prenol lipids |
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P14679 | TYR | Tyrosinase | T97035 | SEA |
| Q16236 | NFE2L2 | Nuclear factor erythroid 2-related factor 2 | T88505 | SEA |
| Q16637 | SMN2 | Survival motor neuron protein | T71907 | SEA |
| O75751 | EMTH | Organic cation transporter 3 | T55948 | SEA |
| Q9C000 | NLRP1 | NLR pyrin domain containing 1 | T86910 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T97035 | DI0007 | Acquired hypermelanosis | [ICD-11: ED60] | P14679 | TYR |
| T97035 | DI0008 | Acquired hypomelanotic disorder | [ICD-11: ED63] | P14679 | TYR |
| T88505 | DI0038 | Ataxic disorder | [ICD-11: 8A03] | Q16236 | NFE2L2 |
| T71907 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | Q16637 | SMN2 |
