Compound details
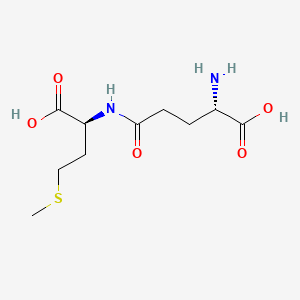
gamma-Glutamylmethionine
| Compound ID | CDAMM00354 |
|---|---|
| Common name | gamma-Glutamylmethionine | IUPAC name | (2S)-2-amino-5-[[(1S)-1-carboxy-3-methylsulfanylpropyl]amino]-5-oxopentanoic acid |
| Molecular formula | C10H18N2O5S |
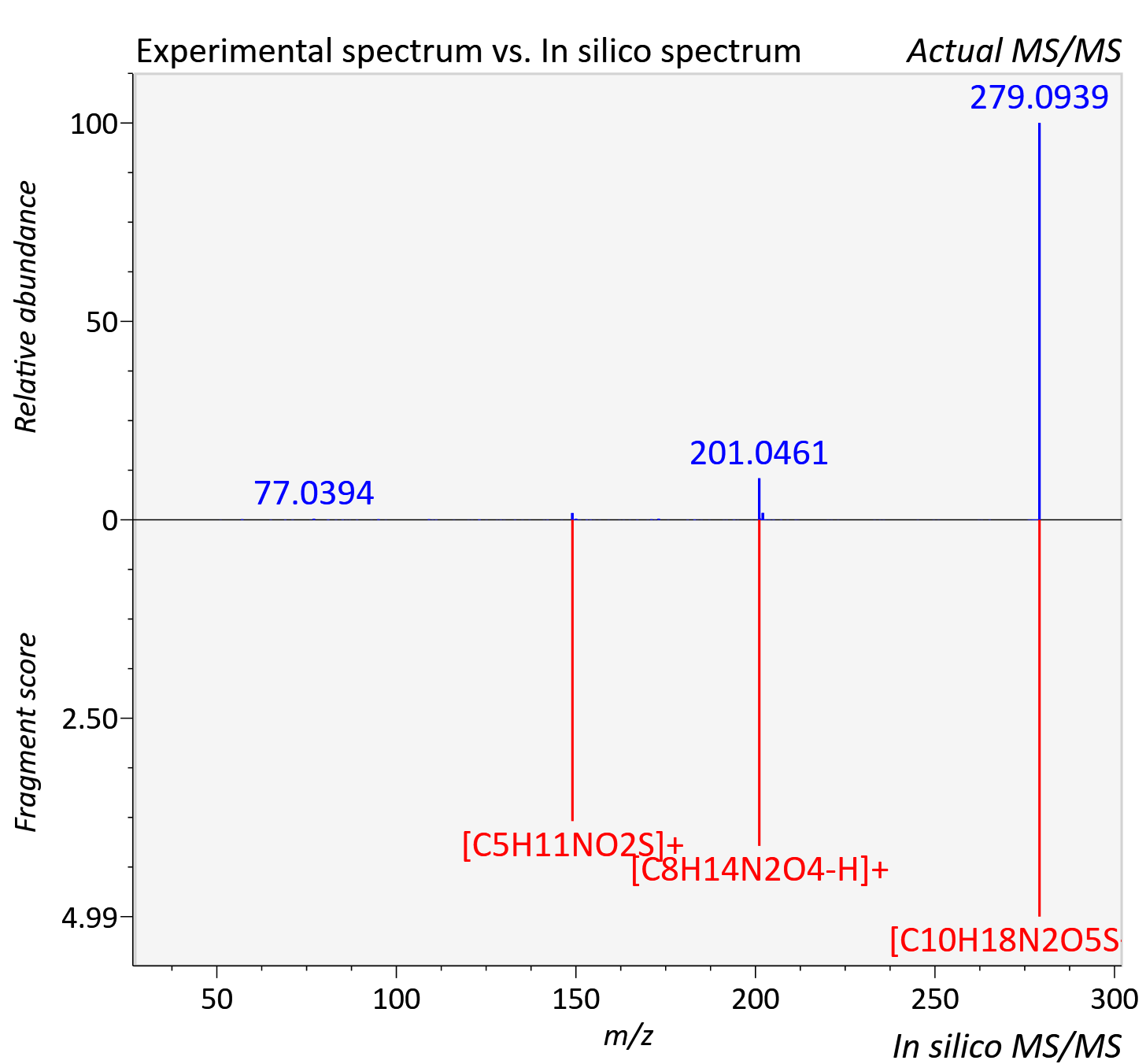
Experimental data
| Retention time | 58.105 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 279.096 | Theoretical mz | 279.101 |
| Error | 13.6 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.42 |
Identifiers and class information
| Inchi key | RQNSKRXMANOPQY-WZTWBHKBNA-N |
|---|---|
| Smiles | O=C(O)C(N)CCC(=O)NC(C(=O)O)CCSC |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SEA |
| P09960 | LTA4H | Leukotriene A4 hydrolase | T03691 | SEA |
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| P43005 | SLC1A1 | Excitatory amino acid transporter 3 | T31721 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| Q13002 | GRIK2 | Glutamate receptor ionotropic kainate 2 | T58178 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| Q07075 | ENPEP | Aminopeptidase A | T31956 | SEA |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SEA |
| P09211 | GSTP1 | Glutathione S-transferase Pi | T21669 | SEA |
| P08263 | GSTA1 | Glutathione S-transferase A1 | T17221 | SEA |
| P04439 | HLA-A | HLA class I histocompatibility antigen A-3 | T08985 | SEA |
| Q16581 | C3AR1 | C3a anaphylatoxin chemotactic receptor | T30219 | SEA |
| P43003 | SLC1A3 | Excitatory amino acid transporter 1 | T86582 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| Q92820 | GGH | Gamma-glutamyl hydrolase | T71646 | SEA |
| P29371 | TACR3 | Neurokinin 3 receptor | T29683 | SEA |
| Q08752 | PPID | Peptidyl-prolyl cis-trans isomerase D | T52447 | SEA |
| P22102 | GART | GAR transformylase | T72042 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| P21462 | FPR1 | Formyl peptide receptor 1 | T87831 | SEA |
| Q04756 | HGFAC | Hepatocyte growth factor activator | T38584 | SEA |
| P09210 | GSTA2 | Glutathione S-transferase A2 | T53510 | SEA |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| P09958 | FUR | HUMAN dibasic-processing enzyme | T98945 | SEA |
| Q9Y4G9 | PCSK6 | Proprotein convertase subtilisin/kexin type 6 | T86503 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| P00480 | OTC | Ornithine transcarbamylase | T52098 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
| P49356 | FNTB | Farnesyl protein transferase | T13127 | SEA |
| Q92887 | CMOAT | Multidrug resistance-associated protein 2 | T61792 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T03691 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P09960 | LTA4H |
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T31956 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q07075 | ENPEP |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
| T21669 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P09211 | GSTP1 |
| T29683 | DI0254 | Menopausal disorder | [ICD-11: GA30] | P29371 | TACR3 |
| T72042 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P22102 | GART |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
| T87831 | DI0333 | Peptic ulcer | [ICD-11: DA61] | P21462 | FPR1 |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
| T98945 | DI0263 | Middle East Respiratory Syndrome | [ICD-11: 1D64] | P09958 | FUR |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T52098 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P00480 | OTC |
| T13127 | DI0342 | Premature ageing appearance | [ICD-11: LD2B] | P49356 | FNTB |
