Compound details
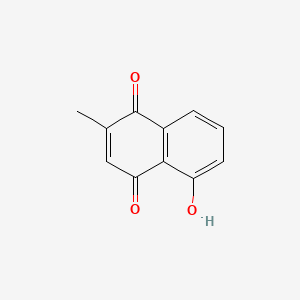
Plumbagin
| Compound ID | CDAMM00350 |
|---|---|
| Common name | Plumbagin | IUPAC name | 5-hydroxy-2-methylnaphthalene-1,4-dione |
| Molecular formula | C11H8O3 |
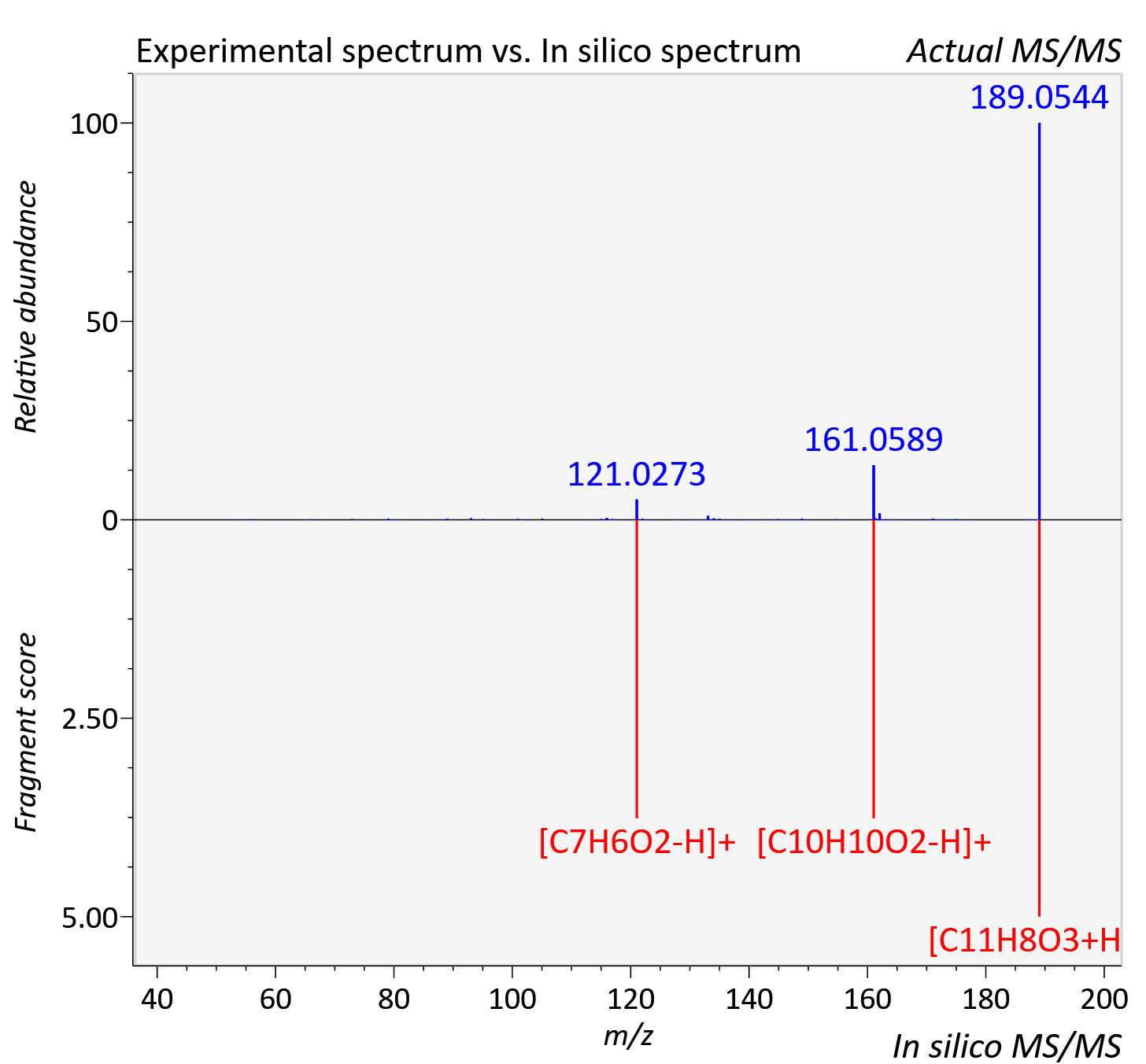
Experimental data
| Retention time | 61.588 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 189.055 | Theoretical mz | 189.055 |
| Error | 1.48 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.26 |
Identifiers and class information
| Inchi key | VCMMXZQDRFWYSE-UHFFFAOYSA-N |
|---|---|
| Smiles | O=C1C=C(C(=O)C=2C=CC=C(O)C12)C |
| Superclass | Benzenoids |
| Class | Naphthalenes |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P14902 | IDO1 | Indoleamine 2,3-dioxygenase | T89697 | SwissTargetPrediction and SEA |
| P08575 | PTPRC | Leukocyte common antigen | T43115 | SEA |
| Q09472 | EP300 | Histone acetyltransferase p300 | T25956 | SwissTargetPrediction and SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SwissTargetPrediction and SEA |
| P04406 | GAPDH | Glyceraldehyde-3-phosphate dehydrogenase liver | T39321 | SEA |
| P00390 | GSR | Glutathione reductase | T30803 | SEA |
| P16885 | PLCG2 | Phospholipase C-gamma-2 | T93922 | SEA |
| P16152 | CBR1 | Carbonyl reductase [NADPH] 1 | T70518 | SEA |
| Q9UBC3 | DNMT3B | DNA (cytosine-5)-methyltransferase 3B | T65501 | SEA |
| O43826 | SLC37A4 | Glucose-6-phosphate translocase | T47306 | SEA |
| O95551 | TDP2 | Tyrosyl-DNA phosphodiesterase 2 | T04696 | SEA |
| P09172 | DBH | Dopamine beta hydroxylase | T74937 | SEA |
| Q9NXA8 | SIRT5 | NAD-dependent deacetylase sirtuin-5 | T91940 | SEA |
| Q15717 | ELAVL1 | ELAV-like protein 1 | T78349 | SEA |
| P02675 | FGB | Fibrin | T19414 | SEA |
| H3BPQ1 | TCF4 | Transcription factor 7-like 2 | T54646 | SEA |
| O75365 | PTP4A3 | Protein tyrosine phosphatase IVA 3 | T78019 | SEA |
| P26447 | S100A4 | S100 calcium-binding protein A4 | T35265 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T89697 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P14902 | IDO1 |
| T43115 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P08575 | PTPRC |
| T43115 | DI0414 | Transplanted organ/tissue | [ICD-11: QB63] | P08575 | PTPRC |
| T25956 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q09472 | EP300 |
| T25956 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q09472 | EP300 |
| T39321 | DI0279 | Muscular atrophy | [ICD-11: 8B61] | P04406 | GAPDH |
| T39321 | DI0280 | Muscular dystrophy | [ICD-11: 8C70] | P04406 | GAPDH |
| T30803 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P00390 | GSR |
| T70518 | DI0037 | Asthma | [ICD-11: CA23] | P16152 | CBR1 |
| T65501 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9UBC3 | DNMT3B |
| T74937 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P09172 | DBH |
| T74937 | DI0341 | Post-traumatic stress disorder | [ICD-11: 6B40] | P09172 | DBH |
| T19414 | DI0052 | Bleeding disorder | [ICD-11: GA20-GA21] | P02675 | FGB |
| T19414 | DI0386 | Skin postprocedural disorder | N.A | P02675 | FGB |
| T35265 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P26447 | S100A4 |
