Compound details
Riboflavin
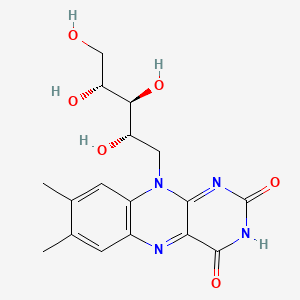
| Compound ID | CDAMM00271 |
|---|---|
| Common name | Riboflavin | IUPAC name | 7,8-dimethyl-10-[(2S,3S,4R)-2,3,4,5-tetrahydroxypentyl]benzo[g]pteridine-2,4-dione |
| Molecular formula | C17H20N4O6 |
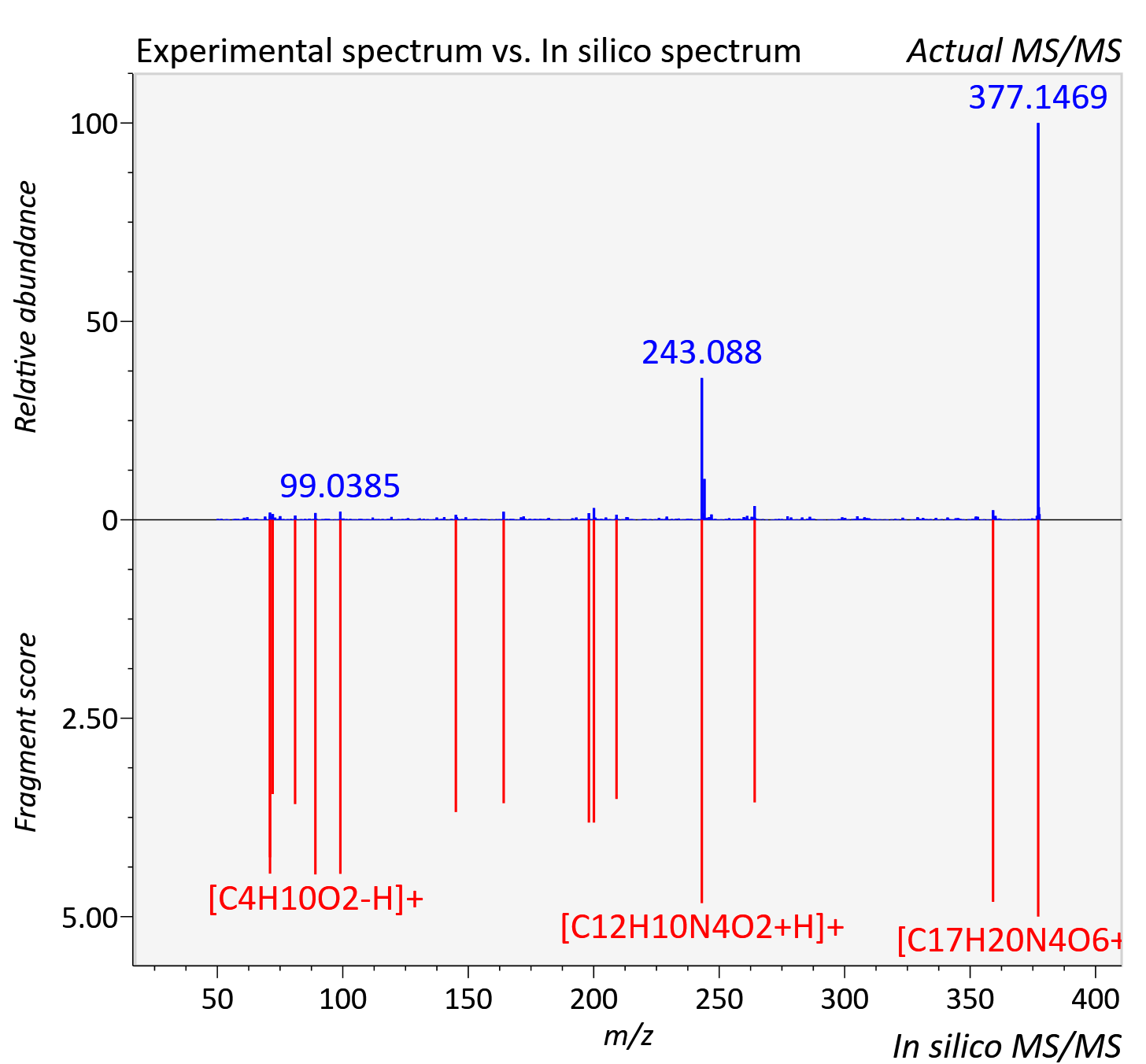
Experimental data
| Retention time | 31.04 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 377.148 | Theoretical mz | 377.146 |
| Error | 6.46 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.05 |
Identifiers and class information
| Inchi key | SFUVCMKSYKHYLD-FNORWQNLSA-N |
|---|---|
| Smiles | CC1=CC2=C(C=C1C)N(C3=NC(=O)NC(=O)C3=N2)CC(C(C(CO)O)O)O |
| Superclass | Phenylpropanoids and polyketides |
| Class | Cinnamic acids and derivatives |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T30803 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P00390 | GSR |
| T85670 | DI0087 | Chronic pain | [ICD-11: MG30] | P01138 | NGF |
| T85670 | DI0163 | General pain disorder | [ICD-11: 8E43] | P01138 | NGF |
| T85670 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P01138 | NGF |
| T85670 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P01138 | NGF |
