Compound details
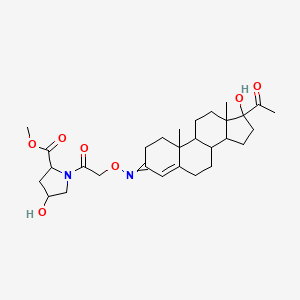
methyl 1-{2-[({1-acetyl-1-hydroxy-9a,11a-dimethyl-1H,2H,3H,3aH,3bH,4H,5H,7H,8H,9H,9aH,9bH,10H,11H,11aH-cyclopenta[a]phenanthren-7-ylidene}amino)oxy]acetyl}-4-hydroxypyrrolidine-2-carboxylate
| Compound ID | CDAMM00214 |
|---|---|
| Common name | methyl 1-{2-[({1-acetyl-1-hydroxy-9a,11a-dimethyl-1H,2H,3H,3aH,3bH,4H,5H,7H,8H,9H,9aH,9bH,10H,11H,11aH-cyclopenta[a]phenanthren-7-ylidene}amino)oxy]acetyl}-4-hydroxypyrrolidine-2-carboxylate | IUPAC name | methyl 1-[2-[(17-acetyl-17-hydroxy-10,13-dimethyl-2,6,7,8,9,11,12,14,15,16-decahydro-1H-cyclopenta[a]phenanthren-3-ylidene)amino]oxyacetyl]-4-hydroxypyrrolidine-2-carboxylate |
| Molecular formula | C29H42N2O7 |
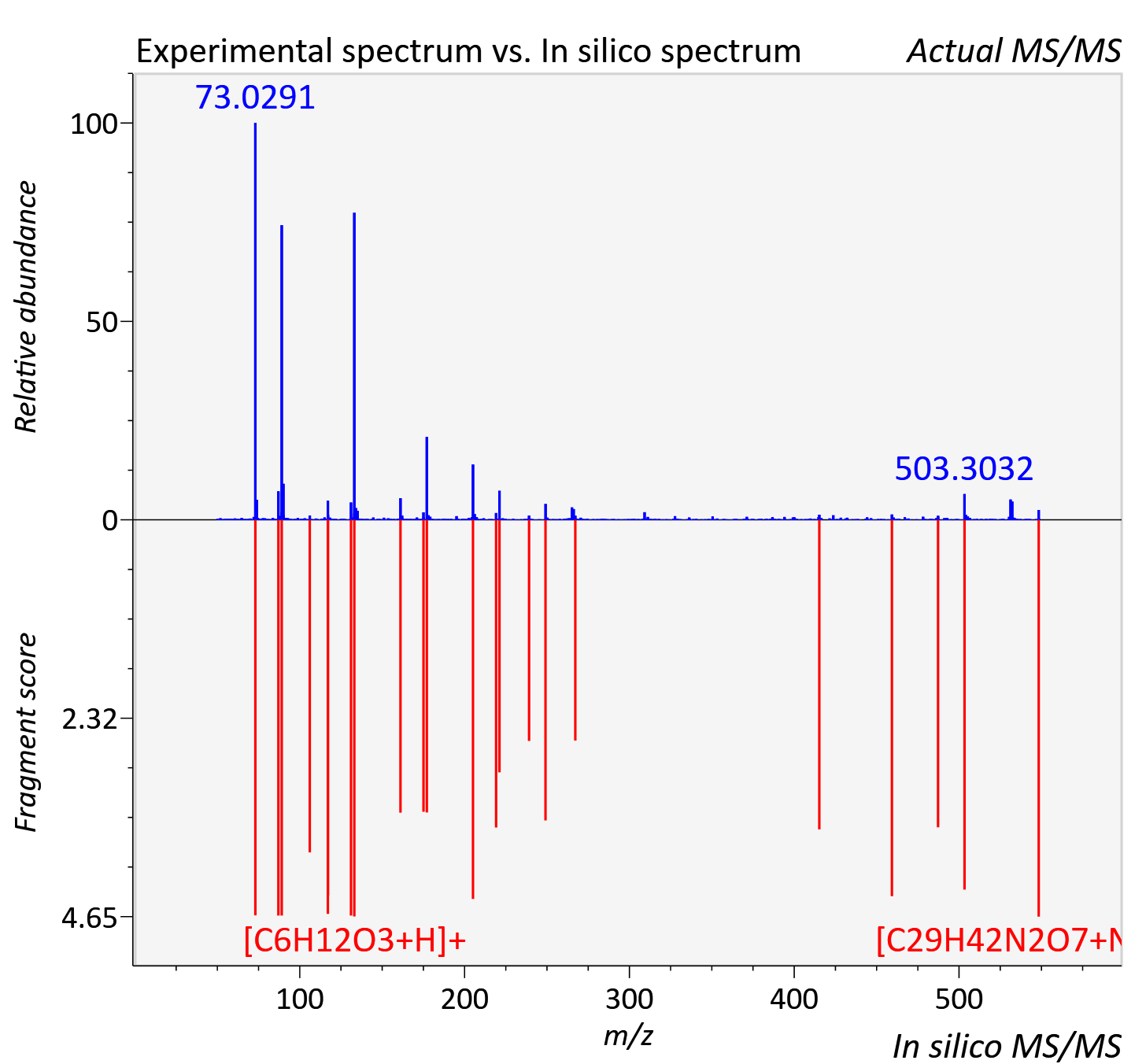
Experimental data
| Retention time | 40.31 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 548.332 | Theoretical mz | 548.333 |
| Error | 1.49 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.5143 |
Identifiers and class information
| Inchi key | CJKRIXOGACCXAJ-UHFFFAOYSA-N |
|---|---|
| Smiles | CC(=O)C1(CCC2C1(CCC3C2CCC4=CC(=NOCC(=O)N5CC(CC5C(=O)OC)O)CCC34C)C)O |
| Superclass | Lipids and lipid-like molecules |
| Class | Steroids and steroid derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08235 | NR3C2 | Mineralocorticoid receptor | T72168 | SEA |
| P04150 | NR3C1 | Glucocorticoid receptor | T40016 | SEA |
| P08185 | SERPINA6 | Corticosteroid binding globulin | T88452 | SEA |
| P11511 | CYP19A1 | Cytochrome P450 19A1 | T13260 | SEA |
| P11413 | G6PD | Glucose-6-phosphate 1-dehydrogenase | T63484 | SEA |
| P05093 | CYP17A1 | Cytochrome P450 17A1 | T89041 | SEA |
| P05231 | IL6 | Interleukin-6 | T32578 | SEA |
| O75751 | EMTH | Organic cation transporter 3 | T55948 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T72168 | DI0018 | Adrenocortical insufficiency | [ICD-11: 5A74] | P08235 | NR3C2 |
| T72168 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P08235 | NR3C2 |
| T72168 | DI0100 | Contraceptive management | [ICD-11: QA21] | P08235 | NR3C2 |
| T72168 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08235 | NR3C2 |
| T72168 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P08235 | NR3C2 |
| T72168 | DI0194 | Hypo-osmolality/hyponatraemia | [ICD-11: 5C72] | P08235 | NR3C2 |
| T40016 | DI0016 | Adaptive immunity immunodeficiency | [ICD-11: 4A01] | P04150 | NR3C1 |
| T40016 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P04150 | NR3C1 |
| T40016 | DI0037 | Asthma | [ICD-11: CA23] | P04150 | NR3C1 |
| T40016 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P04150 | NR3C1 |
| T40016 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P04150 | NR3C1 |
| T40016 | DI0280 | Muscular dystrophy | [ICD-11: 8C70] | P04150 | NR3C1 |
| T40016 | DI0314 | Oesophagitis | [ICD-11: DA24] | P04150 | NR3C1 |
| T40016 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P04150 | NR3C1 |
| T40016 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P04150 | NR3C1 |
| T40016 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P04150 | NR3C1 |
| T40016 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | P04150 | NR3C1 |
| T88452 | DI0037 | Asthma | [ICD-11: CA23] | P08185 | SERPINA6 |
| T88452 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P08185 | SERPINA6 |
| T88452 | DI0362 | Respiratory system disease | [ICD-11: CB40-CB7Z] | P08185 | SERPINA6 |
| T13260 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P11511 | CYP19A1 |
| T13260 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P11511 | CYP19A1 |
| T63484 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P11413 | G6PD |
| T89041 | DI0346 | Prostate cancer | [ICD-11: 2C82] | P05093 | CYP17A1 |
| T32578 | DI0028 | Anemia | [ICD-11: 3A00-3A9Z] | P05231 | IL6 |
