Compound details
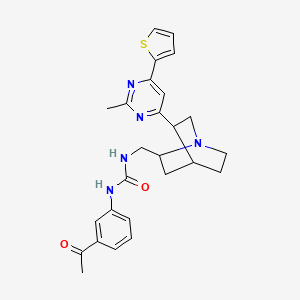
1-(3-acetylphenyl)-3-({5-[2-methyl-6-(thiophen-2-yl)pyrimidin-4-yl]-1-azabicyclo[2.2.2]octan-2-yl}methyl)urea
| Compound ID | CDAMM00193 |
|---|---|
| Common name | 1-(3-acetylphenyl)-3-({5-[2-methyl-6-(thiophen-2-yl)pyrimidin-4-yl]-1-azabicyclo[2.2.2]octan-2-yl}methyl)urea | IUPAC name | 1-(3-acetylphenyl)-3-[[5-(2-methyl-6-thiophen-2-ylpyrimidin-4-yl)-1-azabicyclo[2.2.2]octan-2-yl]methyl]urea |
| Molecular formula | C26H29N5O2S |
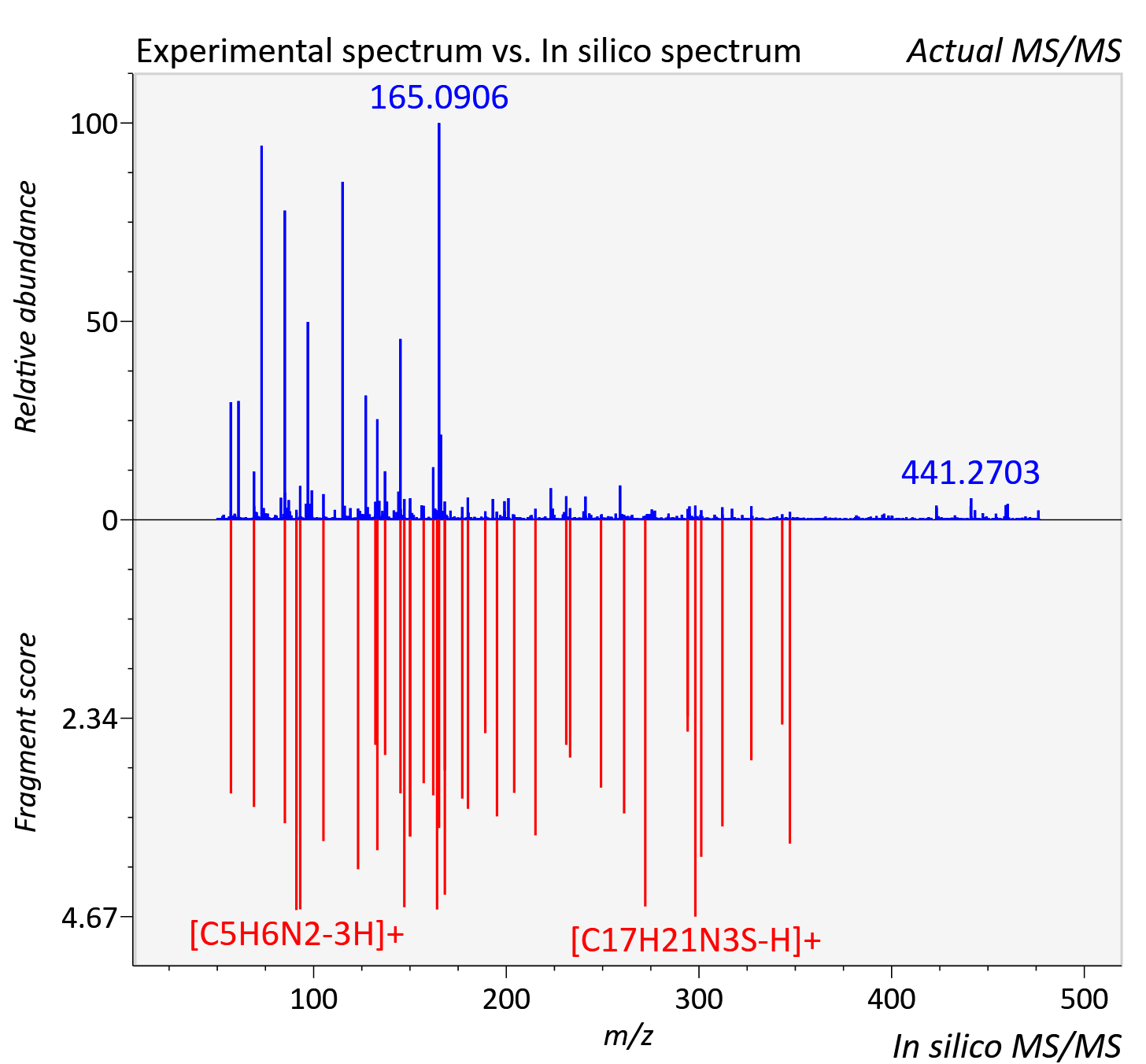
Experimental data
| Retention time | 46.49 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 476.212 | Theoretical mz | 476.211 |
| Error | 1.11 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.7742 |
Identifiers and class information
| Inchi key | VNPRPSZAIQIUCD-UHFFFAOYSA-N |
|---|---|
| Smiles | CC1=NC(=CC(=N1)C2CN3CCC2CC3CNC(=O)NC4=CC=CC(=C4)C(=O)C)C5=CC=CS5 |
| Superclass | Organic oxygen compounds |
| Class | Organooxygen compounds |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P34913 | EPHX2 | Epoxide hydratase | T35734 | SEA |
| P48544 | KCNJ5 | Kir3.1/Kir3.4 | T78692 | SEA |
| P48051 | KCNJ6 | Kir3.1/Kir3.2 | T17721 | SEA |
| Q9UBS0 | RPS6KB2 | Ribosomal protein S6 kinase beta-2 | T42760 | SEA |
| P48549 | KCNJ3 | Inward rectifier potassium channel Kir3.1 | T38012 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T35734 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P34913 | EPHX2 |
| T35734 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P34913 | EPHX2 |
| T78692 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P48544 | KCNJ5 |
| T78692 | DI0183 | Hodgkin lymphoma | [ICD-11: 2B30] | P48544 | KCNJ5 |
| T42760 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q9UBS0 | RPS6KB2 |
