Compound details
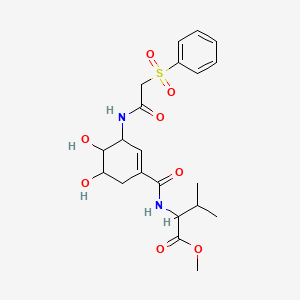
methyl 2-({3-[2-(benzenesulfonyl)acetamido]-4,5-dihydroxycyclohex-1-en-1-yl}formamido)-3-methylbutanoate
| Compound ID | CDAMM00191 |
|---|---|
| Common name | methyl 2-({3-[2-(benzenesulfonyl)acetamido]-4,5-dihydroxycyclohex-1-en-1-yl}formamido)-3-methylbutanoate | IUPAC name | methyl 2-[[3-[[2-(benzenesulfonyl)acetyl]amino]-4,5-dihydroxycyclohexene-1-carbonyl]amino]-3-methylbutanoate |
| Molecular formula | C21H28N2O8S |
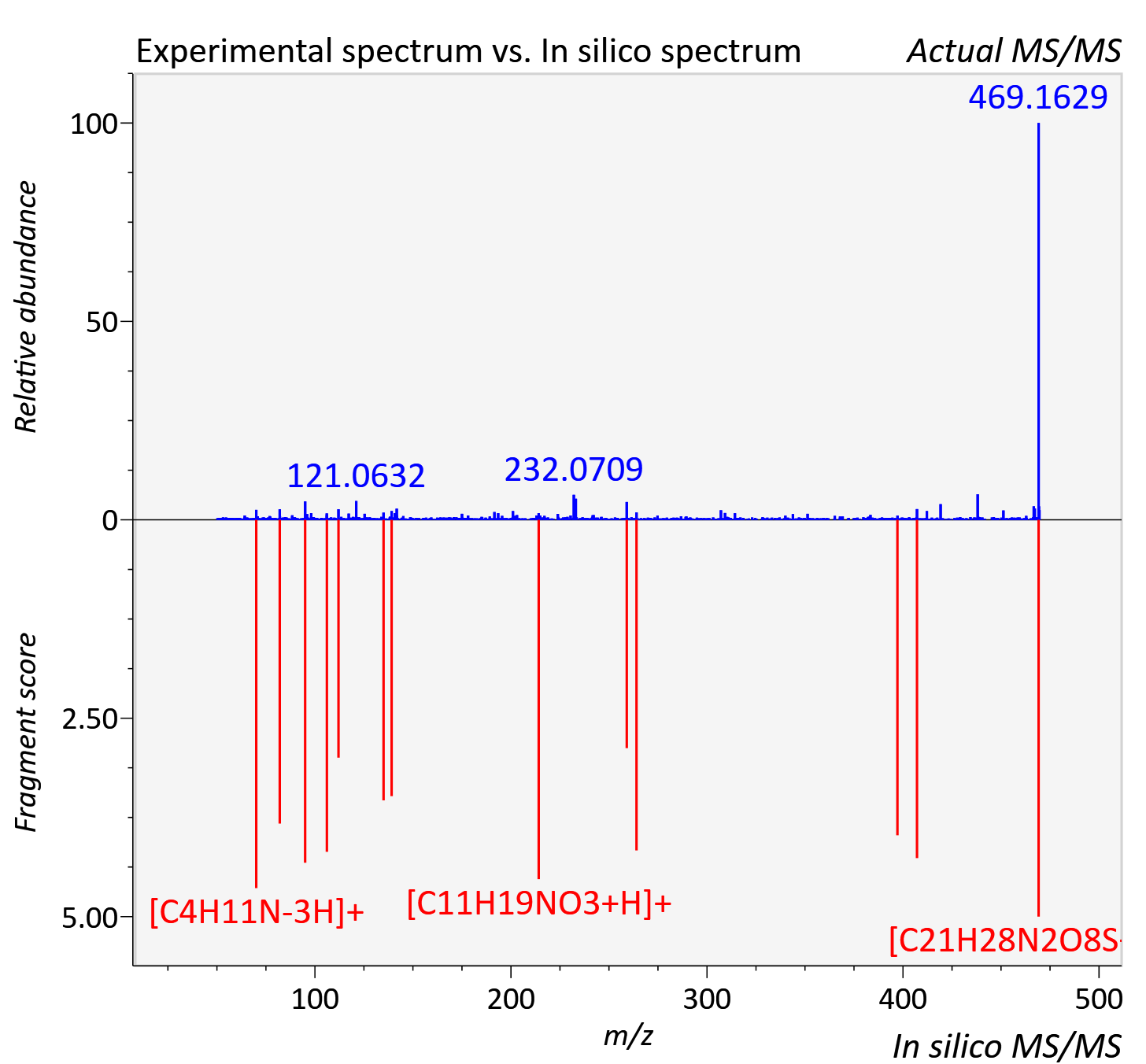
Experimental data
| Retention time | 29.69 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 469.164 | Theoretical mz | 469.164 |
| Error | 0.87 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.1177 |
Identifiers and class information
| Inchi key | FVSLFYHIKZBTGR-UHFFFAOYSA-N |
|---|---|
| Smiles | CC(C)C(C(=O)OC)NC(=O)C1=CC(C(C(C1)O)O)NC(=O)CS(=O)(=O)C2=CC=CC=C2 |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T67102 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P07339 | CTSD |
| T95286 | DI0219 | Ischaemic/haemorrhagic stroke | [ICD-11: 8B20] | P20916 | MAG |
| T41263 | DI0078 | Cholera | [ICD-11: 1A00] | Q7RTX1 | TAS1R1 |
