Compound details
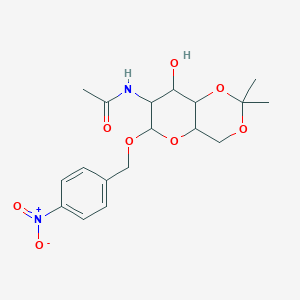
N-{8-hydroxy-2,2-dimethyl-6-[(4-nitrophenyl)methoxy]-hexahydro-2H-pyrano[3,2-d][1,3]dioxin-7-yl}acetamide
| Compound ID | CDAMM00171 |
|---|---|
| Common name | N-{8-hydroxy-2,2-dimethyl-6-[(4-nitrophenyl)methoxy]-hexahydro-2H-pyrano[3,2-d][1,3]dioxin-7-yl}acetamide | IUPAC name | N-[8-hydroxy-2,2-dimethyl-6-[(4-nitrophenyl)methoxy]-4,4a,6,7,8,8a-hexahydropyrano[3,2-d][1,3]dioxin-7-yl]acetamide |
| Molecular formula | C18H24N2O8 |
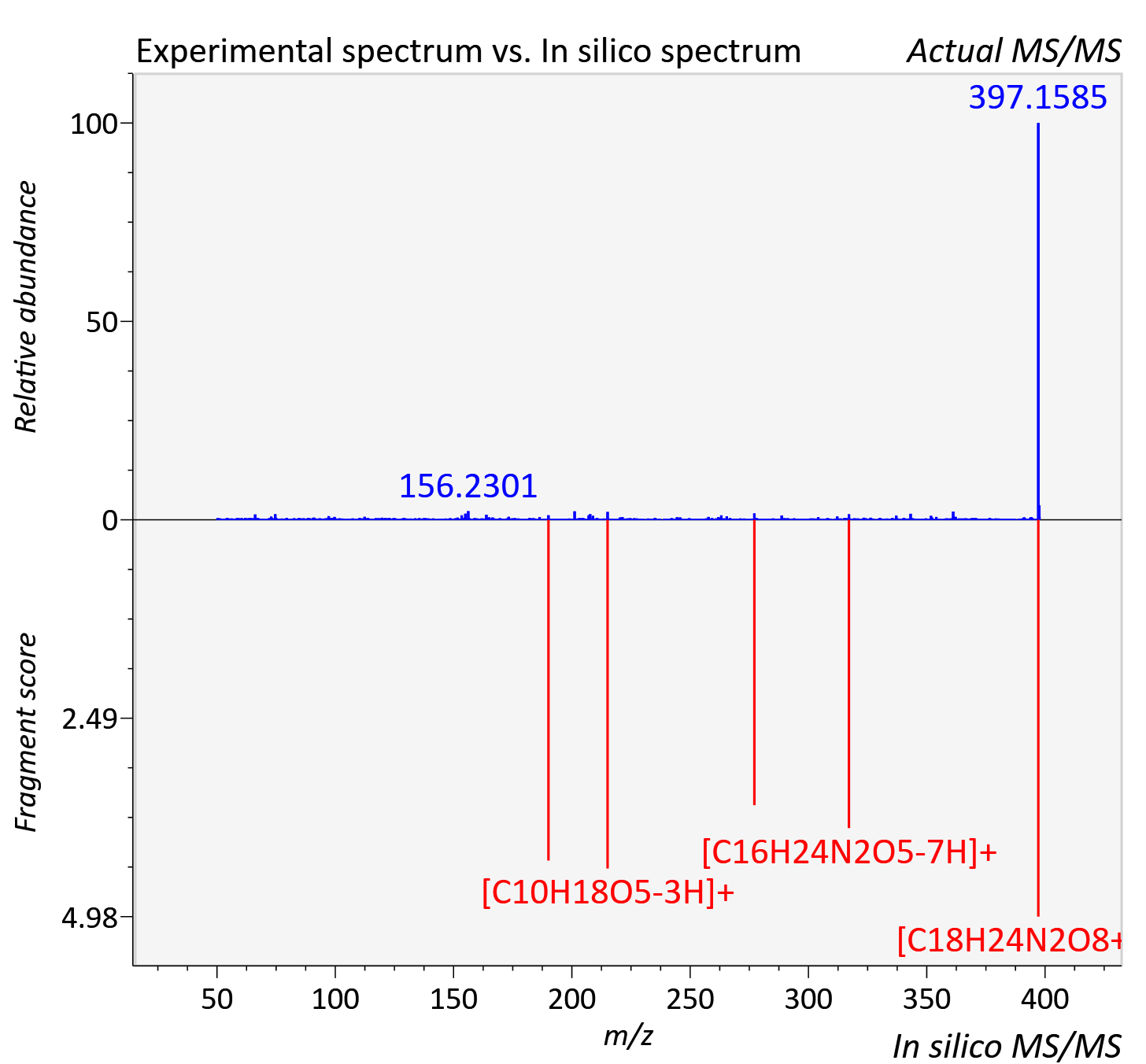
Experimental data
| Retention time | 37.23 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 397.16 | Theoretical mz | 397.16 |
| Error | 1.1 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.8178 |
Identifiers and class information
| Inchi key | OOFFXOCMFXFNIN-UHFFFAOYSA-N |
|---|---|
| Smiles | CC(=O)NC1C(C2C(COC(O2)(C)C)OC1OCC3=CC=C(C=C3)[N+](=O)[O-])O |
| Superclass | Organoheterocyclic compounds |
| Class | Pyranodioxins |
Plant source
Pharmacokinetic properties
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T72038 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | P17931 | LGALS3 |
| T72038 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P17931 | LGALS3 |
| T72038 | DI0252 | Melanoma | [ICD-11: 2C30] | P17931 | LGALS3 |
| T72038 | DI0302 | Non-alcoholic fatty liver disease | [ICD-11: DB92] | P17931 | LGALS3 |
| T72038 | DI0351 | Psoriasis | [ICD-11: EA90] | P17931 | LGALS3 |
| T67894 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | O15455 | TLR3 |
