Compound details
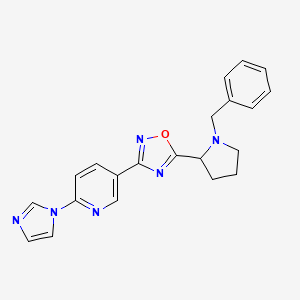
5-[5-(1-benzylpyrrolidin-2-yl)-1,2,4-oxadiazol-3-yl]-2-(1H-imidazol-1-yl)pyridine
| Compound ID | CDAMM00159 |
|---|---|
| Common name | 5-[5-(1-benzylpyrrolidin-2-yl)-1,2,4-oxadiazol-3-yl]-2-(1H-imidazol-1-yl)pyridine | IUPAC name | 5-(1-benzylpyrrolidin-2-yl)-3-(6-imidazol-1-ylpyridin-3-yl)-1,2,4-oxadiazole |
| Molecular formula | C21H20N6O |
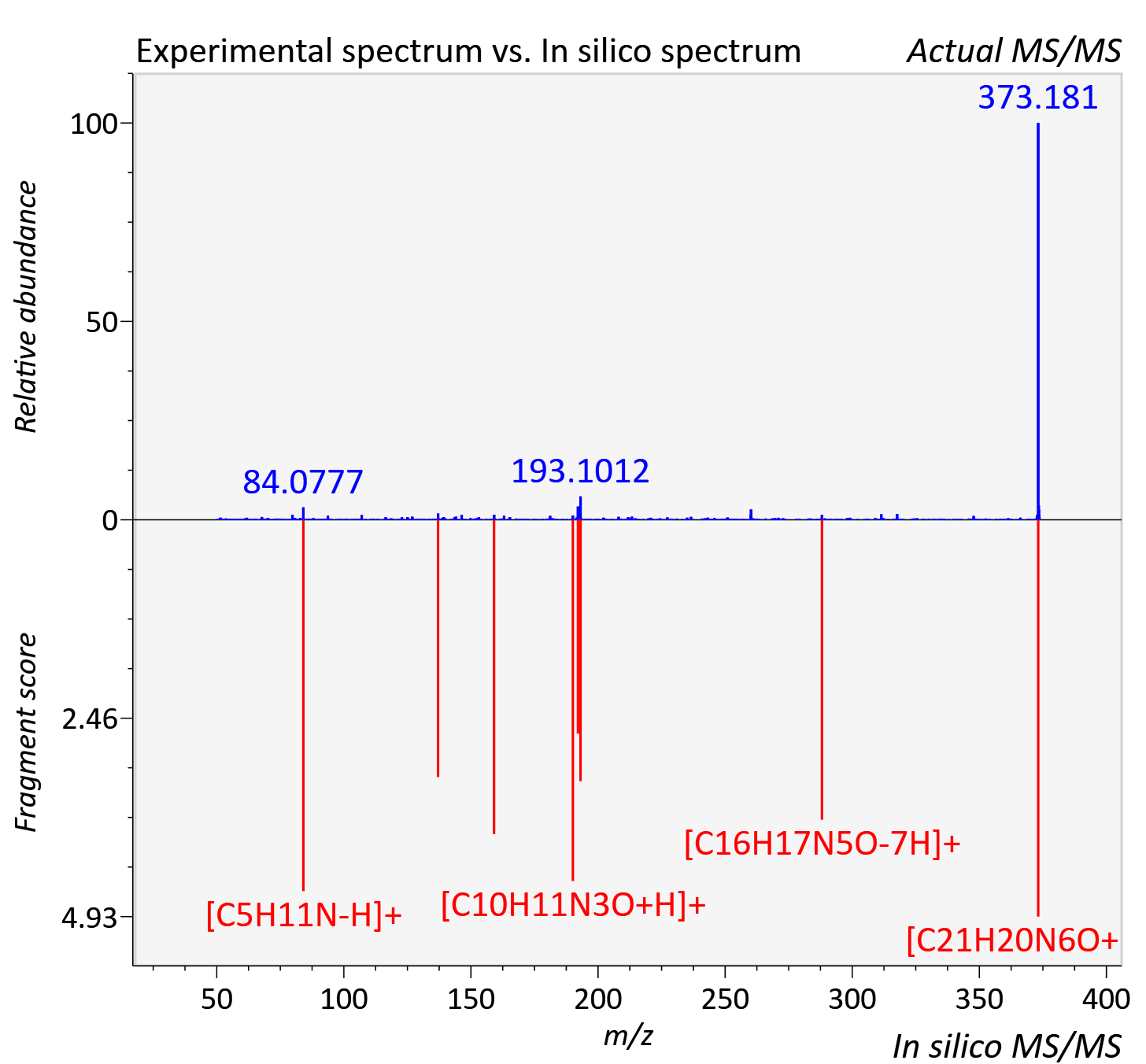
Experimental data
| Retention time | 28.33 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 373.178 | Theoretical mz | 373.177 |
| Error | 1.52 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.3176 |
Identifiers and class information
| Inchi key | CJAPLNIZVVYTIM-UHFFFAOYSA-N |
|---|---|
| Smiles | C1CC(N(C1)CC2=CC=CC=C2)C3=NC(=NO3)C4=CN=C(C=C4)N5C=CN=C5 |
| Superclass | Benzenoids |
| Class | Benzene and substituted derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SEA |
| P41594 | GRM5 | Metabotropic glutamate receptor 5 | T99347 | SEA |
| P42785 | PRCP | Lysosomal Pro-X carboxypeptidase | T20371 | SEA |
| P55263 | ADK | Adenosine kinase | T91661 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T99347 | DI0152 | Fragile X chromosome | [ICD-11: LD55] | P41594 | GRM5 |
| T99347 | DI0331 | Parkinsonism | [ICD-11: 8A00] | P41594 | GRM5 |
| T91661 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P55263 | ADK |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
