Compound details
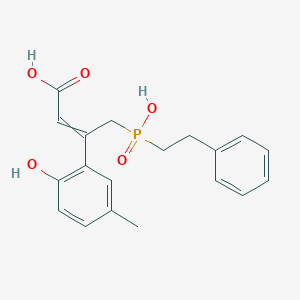
4-[hydroxy(2-phenylethyl)phosphoryl]-3-(2-hydroxy-5-methylphenyl)but-2-enoic acid
| Compound ID | CDAMM00153 |
|---|---|
| Common name | 4-[hydroxy(2-phenylethyl)phosphoryl]-3-(2-hydroxy-5-methylphenyl)but-2-enoic acid | IUPAC name | 3-(2-hydroxy-5-methylphenyl)-4-[hydroxy(2-phenylethyl)phosphoryl]but-2-enoic acid |
| Molecular formula | C19H21O5P |
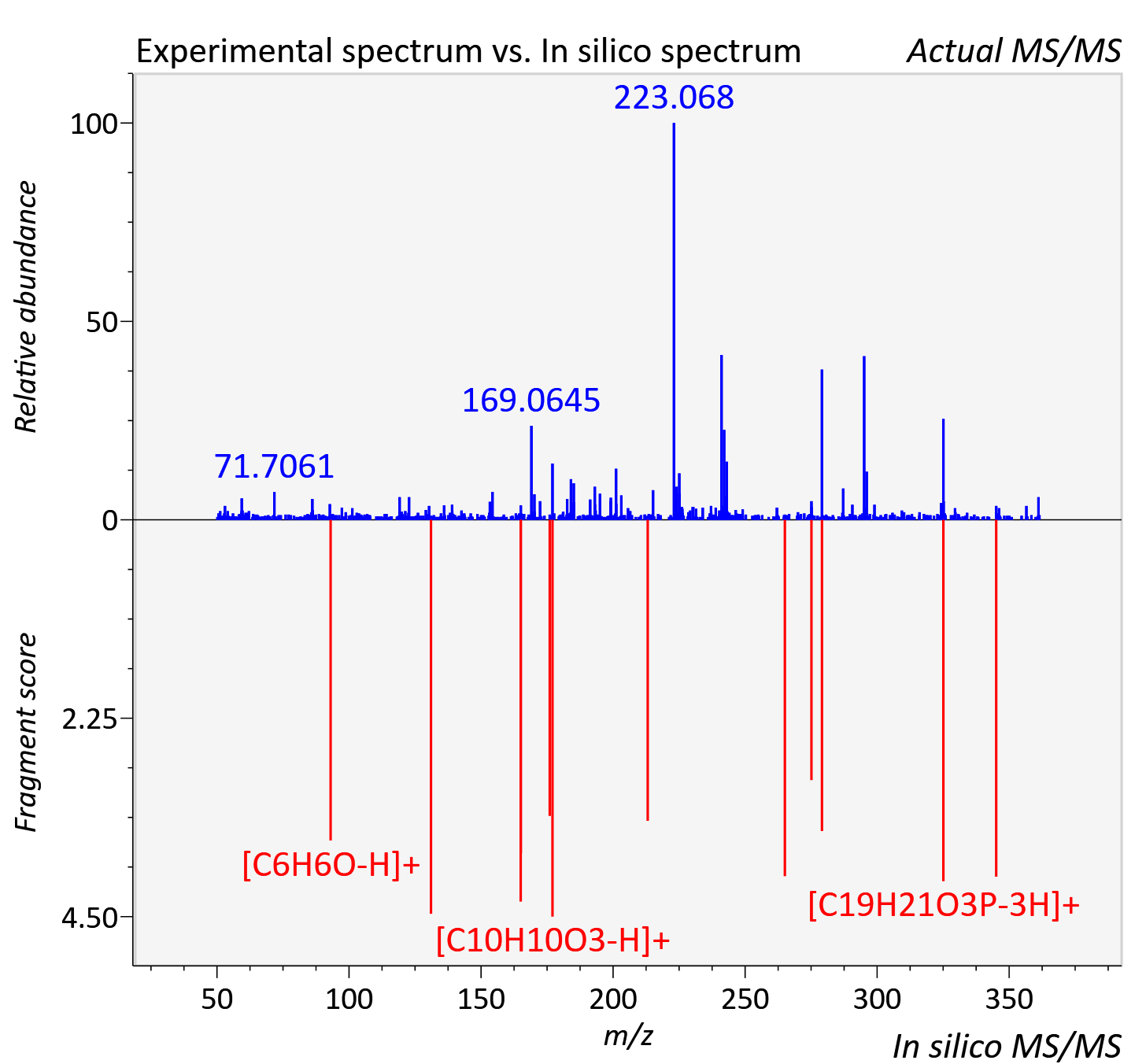
Experimental data
| Retention time | 15.02 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 361.12 | Theoretical mz | 361.12 |
| Error | 0.47 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.3826 |
Identifiers and class information
| Inchi key | DTXBUZQCAJVSQP-UHFFFAOYSA-N |
|---|---|
| Smiles | CC1=CC(=C(C=C1)O)C(=CC(=O)O)CP(=O)(CCC2=CC=CC=C2)O |
| Superclass | Phenylpropanoids and polyketides |
| Class | Cinnamic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P15144 | ANPEP | Aminopeptidase N | T67272 | SEA |
| P16050 | ALOX15 | Arachidonate 15-lipoxygenase | T16042 | SEA |
| Q12791 | KCNMA1 | Calcium-activated potassium channel subunit alpha-1 | T66538 | SEA |
| Q16548 | BCL2A1 | Bcl-2-related protein A1 (by homology) | T12797 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T67272 | DI0351 | Psoriasis | [ICD-11: EA90] | P15144 | ANPEP |
| T66538 | DI0037 | Asthma | [ICD-11: CA23] | Q12791 | KCNMA1 |
| T66538 | DI0154 | Functional bladder disorder | [ICD-11: GC50] | Q12791 | KCNMA1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
