Compound details
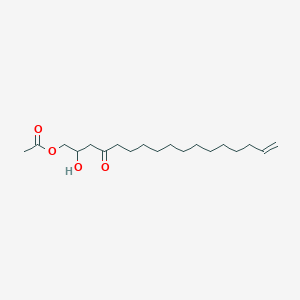
1-Acetoxy-2-hydroxy-16-heptadecen-4-one
| Compound ID | CDAMM00141 |
|---|---|
| Common name | 1-Acetoxy-2-hydroxy-16-heptadecen-4-one | IUPAC name | (2-hydroxy-4-oxoheptadec-16-enyl) acetate |
| Molecular formula | C19H34O4 |
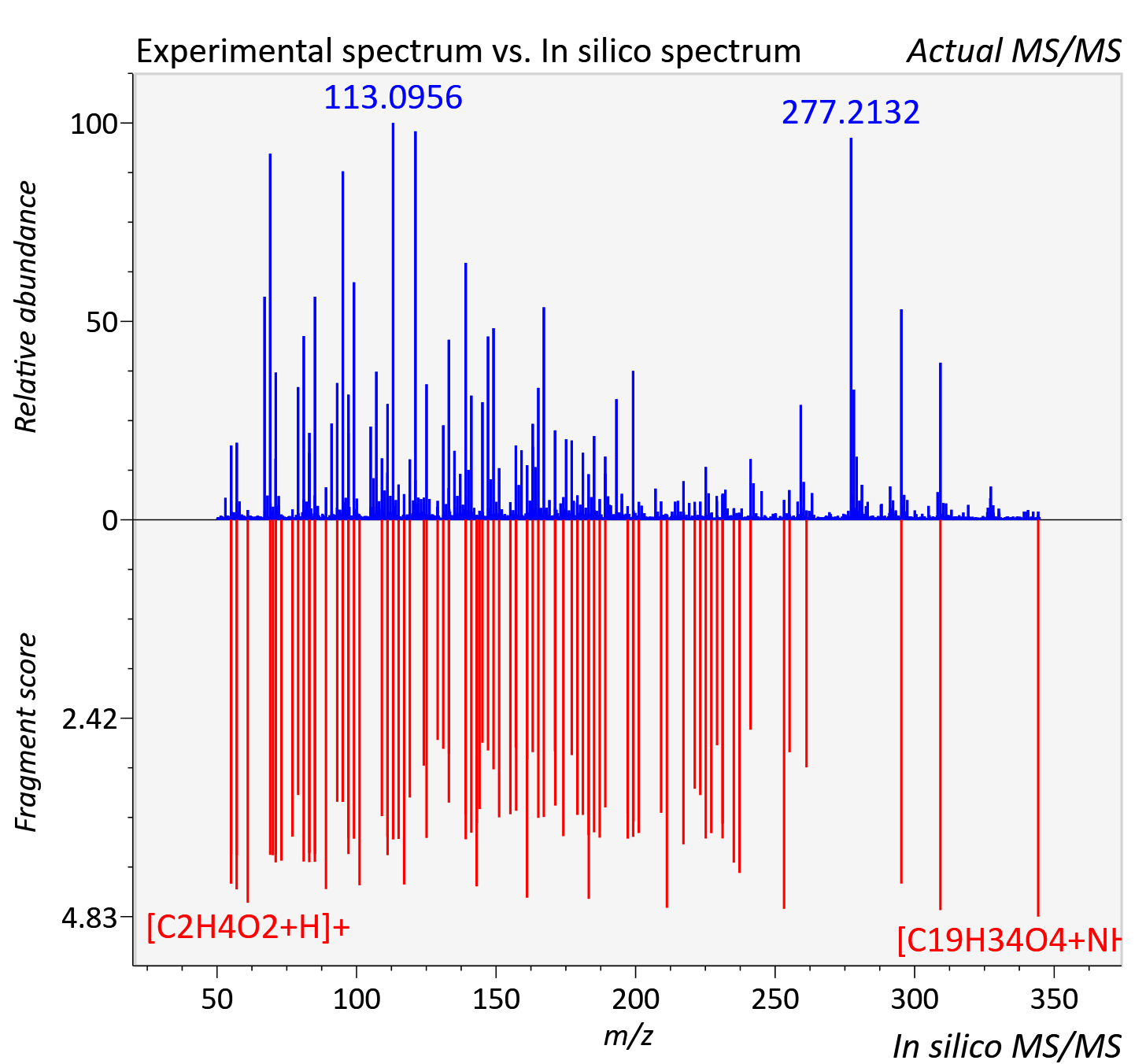
Experimental data
| Retention time | 57.46 |
|---|---|
| Adduct | [M+NH4]+ |
| Actual mz | 344.279 | Theoretical mz | 344.28 |
| Error | 0.95 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.1545 |
Identifiers and class information
| Inchi key | MSSXBCMVFDVFJB-UHFFFAOYNA-N |
|---|---|
| Smiles | CC(=O)OCC(CC(=O)CCCCCCCCCCCC=C)O |
| Superclass | Lipids and lipid-like molecules |
| Class | Fatty Acyls |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| Q9H1C0 | LPAR5 | Lysophosphatidic acid receptor 5 | T18616 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
