Compound details
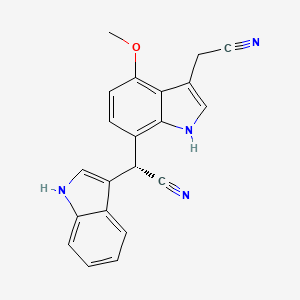
(2S)-2-[3-(Cyanomethyl)-4-Methoxy-1H-Indol-7-Yl]-2-(1H-Indol-3-Yl)Acetonitrile
| Compound ID | CDAMM00136 |
|---|---|
| Common name | (2S)-2-[3-(Cyanomethyl)-4-Methoxy-1H-Indol-7-Yl]-2-(1H-Indol-3-Yl)Acetonitrile | IUPAC name | (2R)-2-[3-(cyanomethyl)-4-methoxy-1H-indol-7-yl]-2-(1H-indol-3-yl)acetonitrile |
| Molecular formula | C21H16N4O |
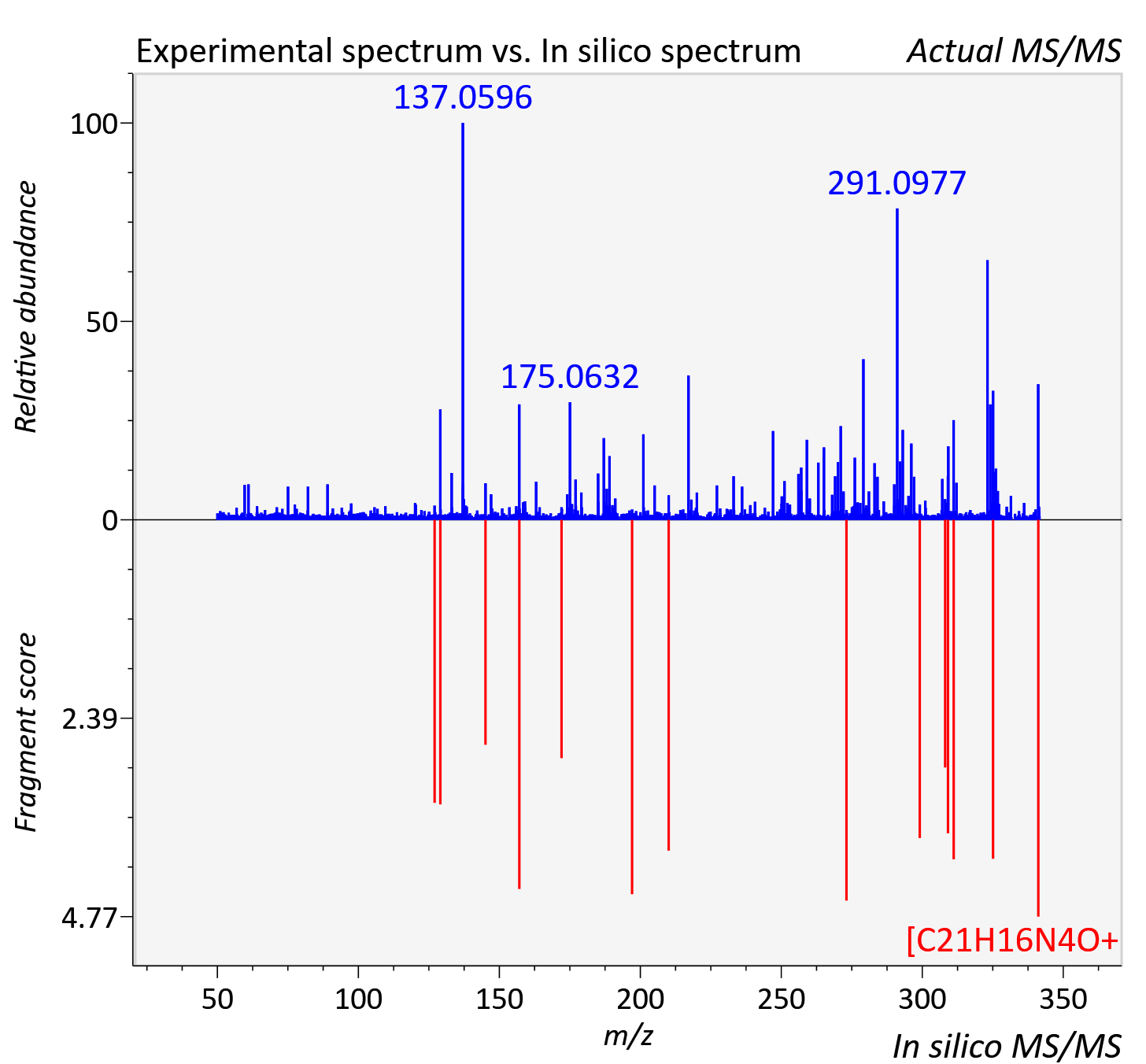
Experimental data
| Retention time | 40.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 341.141 | Theoretical mz | 341.14 |
| Error | 2.69 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.0345 |
Identifiers and class information
| Inchi key | VFCSQPLDMATYSC-INIZCTEOSA-N |
|---|---|
| Smiles | COC1=C2C(=CNC2=C(C=C1)C(C#N)C3=CNC4=CC=CC=C43)CC#N |
| Superclass | Organoheterocyclic compounds |
| Class | Indoles and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q9UQL6 | HDAC5 | Histone deacetylase 5 | T07930 | SEA |
| Q9UKV0 | HDAC9 | Histone deacetylase 9 | T03687 | SEA |
| P35270 | SPR | Substance-P receptor | T47094 | SEA |
| P51511 | MMP15 | Matrix metalloproteinase 15 | T81658 | SEA |
| Q96DB2 | HDAC11 | Histone deacetylase 11 | T97903 | SEA |
| P28336 | NMBR | Neuromedin B receptor | T68887 | SEA |
| P23284 | PPIB | Cyclophilin B | T25847 | SEA |
| Q9ULZ9 | MMP17 | Matrix metalloproteinase-17 | T55616 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T47094 | DI0117 | Depression | [ICD-11: 6A70-6A7Z] | P35270 | SPR |
| T47094 | DI0293 | Nausea/vomiting | [ICD-11: MD90] | P35270 | SPR |
| T68887 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P28336 | NMBR |
| T68887 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P28336 | NMBR |
| T25847 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | P23284 | PPIB |
