Compound details
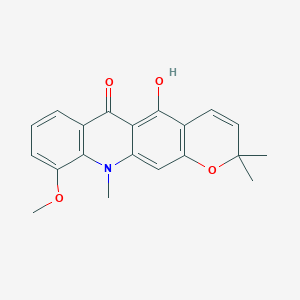
Junosidine
| Compound ID | CDAMM00135 |
|---|---|
| Common name | Junosidine | IUPAC name | 5-hydroxy-10-methoxy-2,2,11-trimethylpyrano[3,2-b]acridin-6-one |
| Molecular formula | C20H19NO4 |
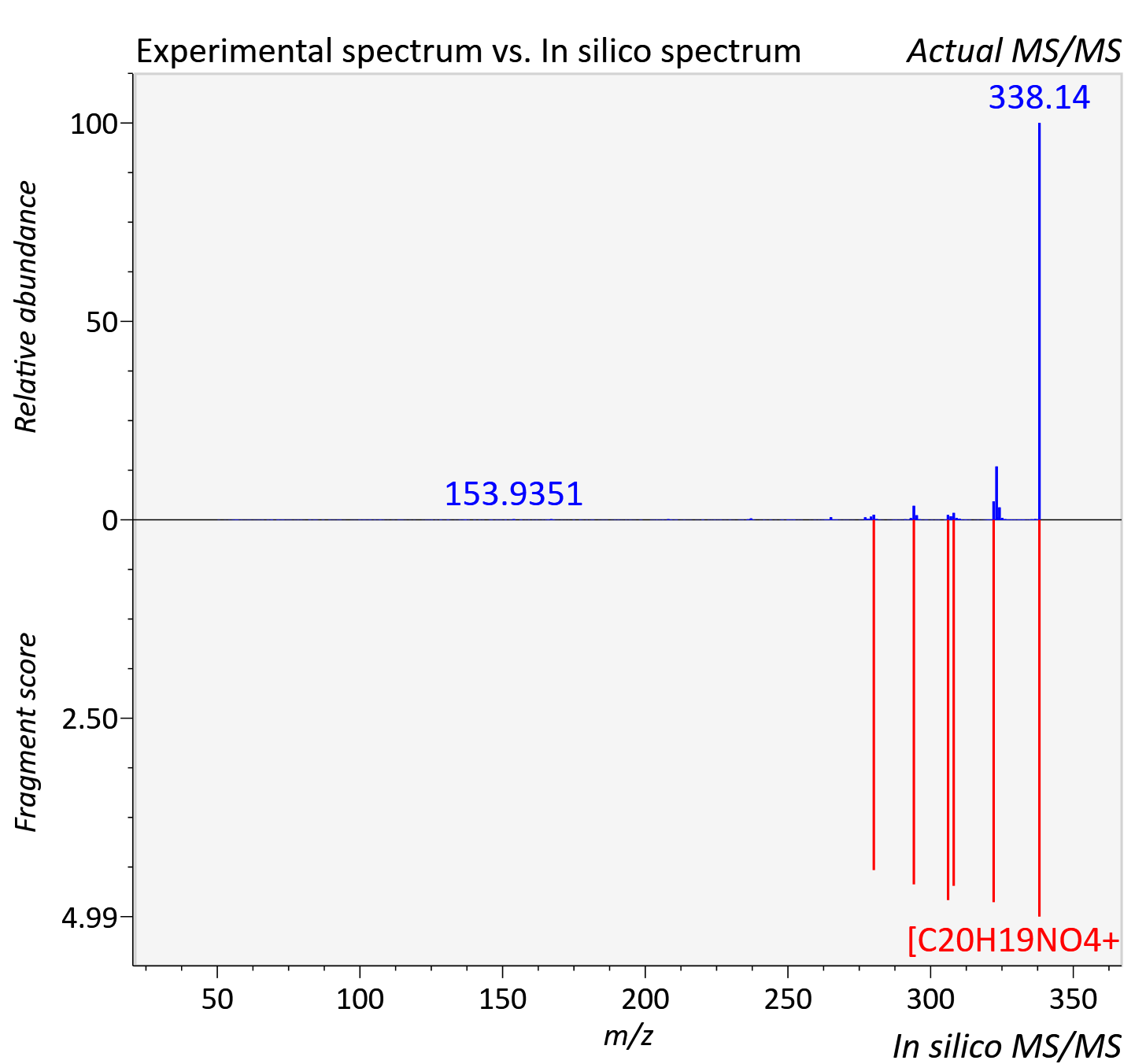
Experimental data
| Retention time | 45.29 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 338.139 | Theoretical mz | 338.139 |
| Error | 0.23 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 7.1671 |
Identifiers and class information
| Inchi key | AUWXWUDANKEVNS-UHFFFAOYSA-N |
|---|---|
| Smiles | CC1(C=CC2=C(O1)C=C3C(=C2O)C(=O)C4=C(N3C)C(=CC=C4)OC)C |
| Superclass | Organoheterocyclic compounds |
| Class | Quinolines and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| O00206 | TLR4 | Toll-like receptor 4 (by homology) | T81443 | SEA |
| Q16678 | CYP1B1 | Cytochrome P450 1B1 | T92521 | SEA |
| O60911 | CTSV | Cathepsin (V and K) | T93653 | SEA |
| Q92887 | CMOAT | Multidrug resistance-associated protein 2 | T61792 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T81443 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | O00206 | TLR4 |
| T81443 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O00206 | TLR4 |
| T81443 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | O00206 | TLR4 |
| T93653 | DI0055 | Bone cancer | [ICD-11: 2B5Z] | O60911 | CTSV |
| T93653 | DI0087 | Chronic pain | [ICD-11: MG30] | O60911 | CTSV |
