Compound details
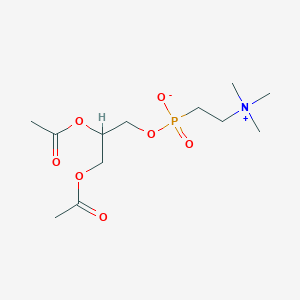
diacylglycerol 2-trimethylaminoethylphosphonate
| Compound ID | CDAMM00128 |
|---|---|
| Common name | diacylglycerol 2-trimethylaminoethylphosphonate | IUPAC name | 2,3-diacetyloxypropoxy-[2-(trimethylazaniumyl)ethyl]phosphinate |
| Molecular formula | C12H24NO7P |
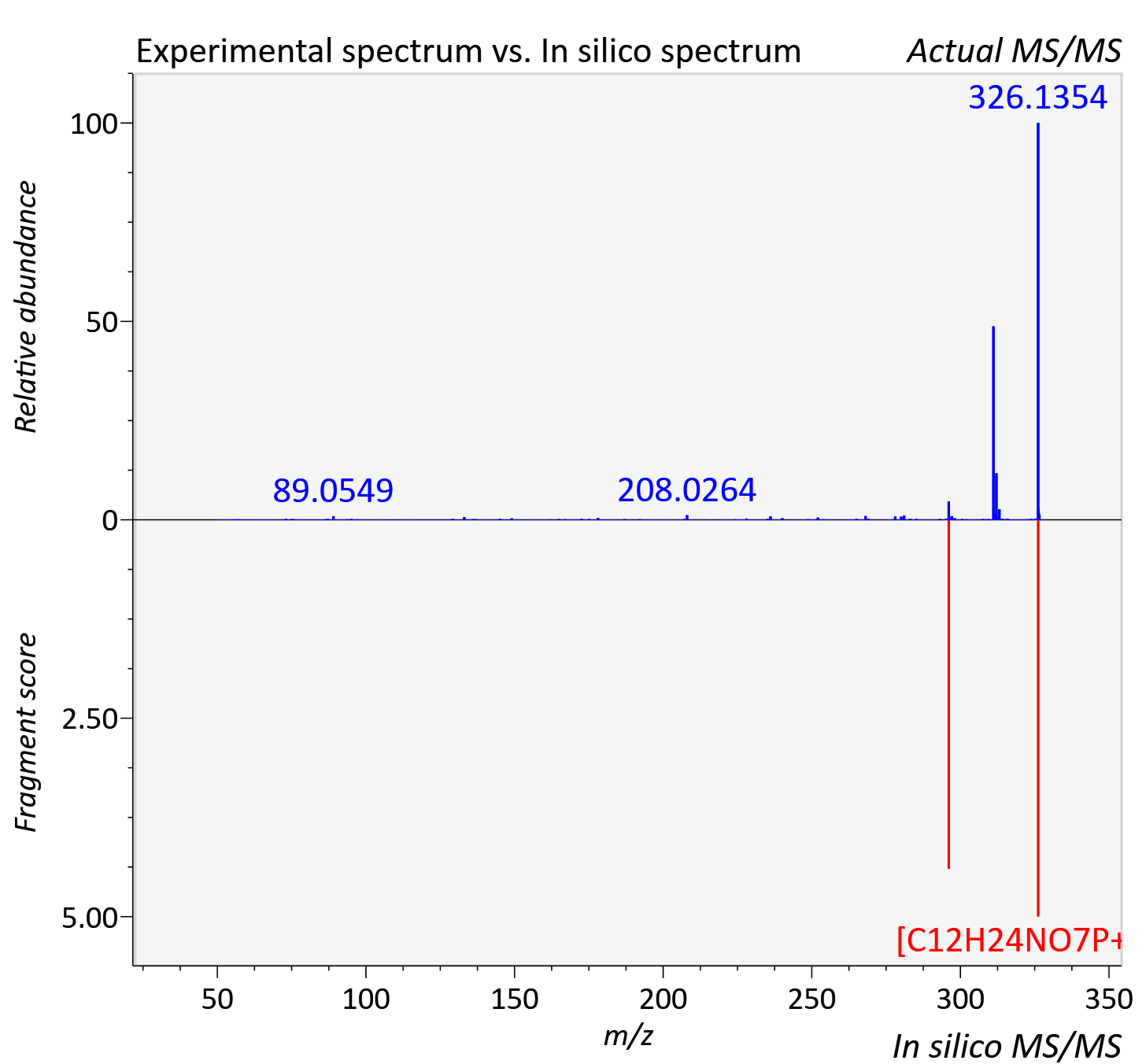
Experimental data
| Retention time | 30.33 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 326.137 | Theoretical mz | 326.136 |
| Error | 3.03 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.8493 |
Identifiers and class information
| Inchi key | SADLPDQPNWOMMM-UHFFFAOYNA-N |
|---|---|
| Smiles | CC(=O)OCC(COP(=O)(CC[N+](C)(C)C)[O-])OC(=O)C |
| Superclass | Lipids and lipid-like molecules |
| Class | Glycerophosphonolipids |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T31514 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | P10253 | GAA |
| T76497 | DI0337 | Pituitary gland disorder | [ICD-11: 5A60-5A61] | P52209 | PGD |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
