Compound details
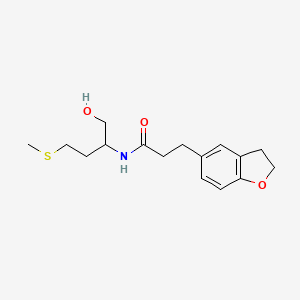
3-(2,3-dihydro-1-benzofuran-5-yl)-N-[1-hydroxy-4-(methylsulfanyl)butan-2-yl]propenamide
| Compound ID | CDAMM00123 |
|---|---|
| Common name | 3-(2,3-dihydro-1-benzofuran-5-yl)-N-[1-hydroxy-4-(methylsulfanyl)butan-2-yl]propenamide | IUPAC name | 3-(2,3-dihydro-1-benzofuran-5-yl)-N-(1-hydroxy-4-methylsulfanylbutan-2-yl)propanamide |
| Molecular formula | C16H23NO3S |
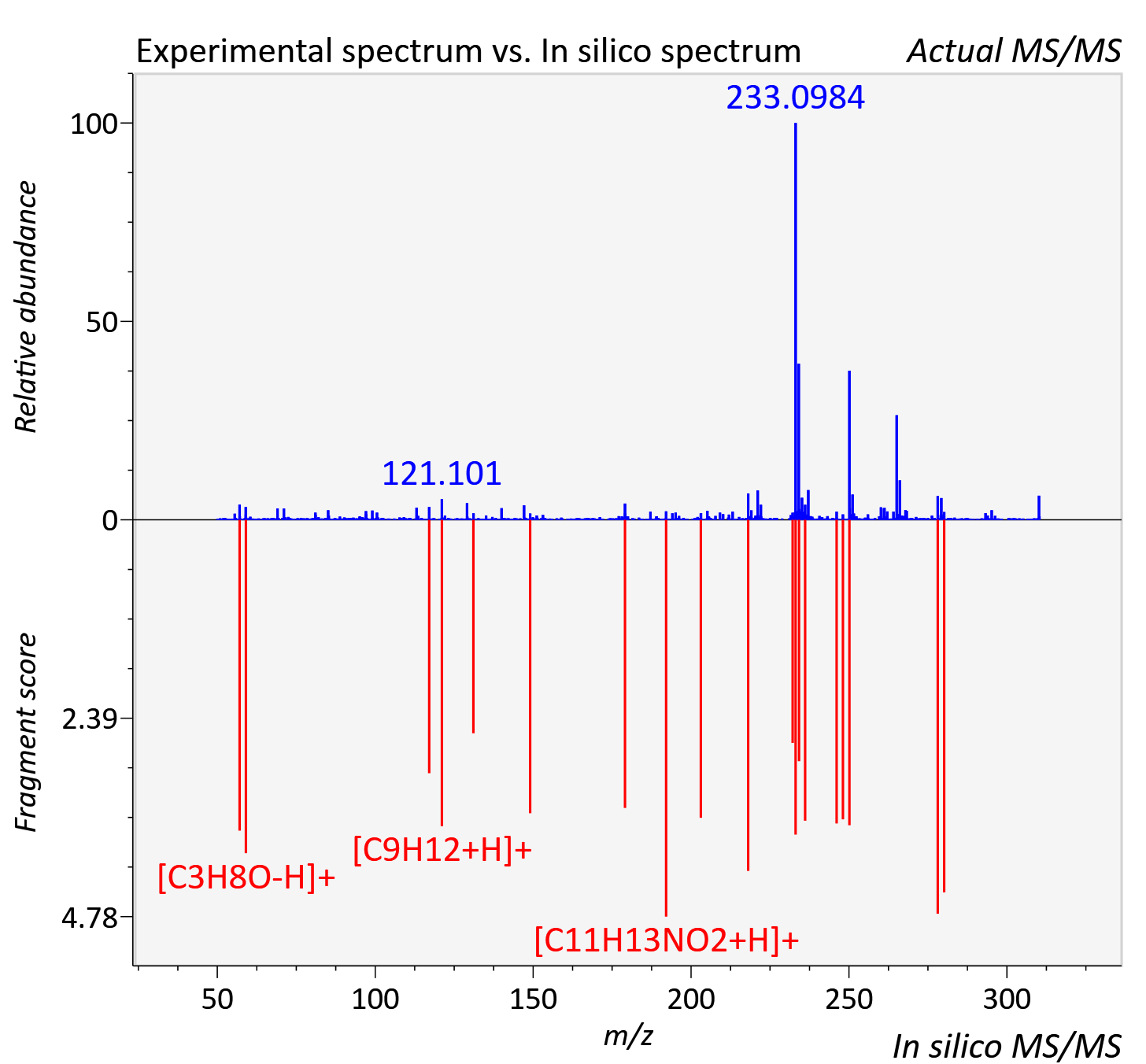
Experimental data
| Retention time | 59.96 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 310.147 | Theoretical mz | 310.147 |
| Error | 1.09 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 4.8389 |
Identifiers and class information
| Inchi key | ORLUDQBZHLUNIF-UHFFFAOYSA-N |
|---|---|
| Smiles | CSCCC(CO)NC(=O)CCC1=CC2=C(C=C1)OCC2 |
| Superclass | Organoheterocyclic compounds |
| Class | Coumarans |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| Q16739 | UGCG | Ceramide glucosyltransferase | T14908 | SEA |
| P22894 | MMP8 | Matrix metalloproteinase 8 | T08856 | SEA |
| P45452 | MMP13 | Matrix metalloproteinase 13 | T34296 | SEA |
| P23946 | CMA1 | Chymase | T05114 | SEA |
| P48039 | MTNR1A | Melatonin receptor 1A | T97613 | SEA |
| P49286 | MTNR1B | Melatonin receptor 1B | T48268 | SEA |
| P39900 | MMP12 | Matrix metalloproteinase 12 | T03500 | SEA |
| P09238 | MMP10 | Matrix metalloproteinase 10 | T40488 | SEA |
| Q9UKP6 | UTS2R | Urotensin II receptor | T49072 | SEA |
| Q08752 | PPID | Peptidyl-prolyl cis-trans isomerase D | T52447 | SEA |
| P49682 | CXCR3 | C-X-C chemokine receptor type 3 | T25315 | SEA |
| P04632 | CAPNS1 | HUMAN calpain-1 or calpain small subunit 1 heterodimer | T00099 | SEA |
| Q7RTX1 | TAS1R1 | Taste receptor | T41263 | SEA |
| P02778 | CXCL10 | C-X-C motif chemokine 10 | T30635 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T14908 | DI0242 | Lysosomal disease | [ICD-11: 5C56] | Q16739 | UGCG |
| T14908 | DI0257 | Metabolic disorder | [ICD-11: 5C50-5D2Z] | Q16739 | UGCG |
| T08856 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P22894 | MMP8 |
| T34296 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P45452 | MMP13 |
| T05114 | DI0039 | Atopic eczema | [ICD-11: EA80] | P23946 | CMA1 |
| T05114 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P23946 | CMA1 |
| T05114 | DI0169 | Gram-positive bacterial infection | [ICD-11: 1B74-1F40] | P23946 | CMA1 |
| T05114 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P23946 | CMA1 |
| T05114 | DI0229 | Left ventricular failure | [ICD-11: BD11] | P23946 | CMA1 |
| T05114 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P23946 | CMA1 |
| T97613 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P48039 | MTNR1A |
| T48268 | DI0214 | Insomnia | [ICD-11: 7A00-7A0Z] | P49286 | MTNR1B |
| T03500 | DI0037 | Asthma | [ICD-11: CA23] | P39900 | MMP12 |
| T03500 | DI0238 | Lung cancer | [ICD-11: 2C25] | P39900 | MMP12 |
| T03500 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P39900 | MMP12 |
| T49072 | DI0037 | Asthma | [ICD-11: CA23] | Q9UKP6 | UTS2R |
| T25315 | DI0178 | Hepatic fibrosis/cirrhosis | [ICD-11: DB93] | P49682 | CXCR3 |
| T25315 | DI0339 | Postoperative inflammation | [ICD-11: 1A00-CA43] | P49682 | CXCR3 |
| T25315 | DI0351 | Psoriasis | [ICD-11: EA90] | P49682 | CXCR3 |
| T25315 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P49682 | CXCR3 |
| T00099 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P04632 | CAPNS1 |
| T41263 | DI0078 | Cholera | [ICD-11: 1A00] | Q7RTX1 | TAS1R1 |
| T30635 | DI0107 | Crohn disease | [ICD-11: DD70] | P02778 | CXCL10 |
| T30635 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P02778 | CXCL10 |
| T30635 | DI0200 | Immune system disease | [ICD-11: 4A01-4B41] | P02778 | CXCL10 |
| T30635 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P02778 | CXCL10 |
| T30635 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P02778 | CXCL10 |
| T30635 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P02778 | CXCL10 |
