Compound details
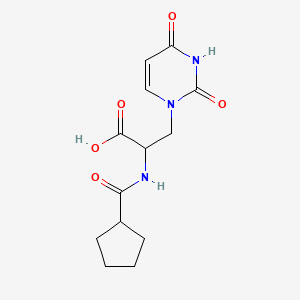
2-(cyclopentylformamido)-3-(2,4-dioxo-1,2,3,4-tetrahydropyrimidin-1-yl)propanoic acid
| Compound ID | CDAMM00112 |
|---|---|
| Common name | 2-(cyclopentylformamido)-3-(2,4-dioxo-1,2,3,4-tetrahydropyrimidin-1-yl)propanoic acid | IUPAC name | 2-(cyclopentanecarbonylamino)-3-(2,4-dioxopyrimidin-1-yl)propanoic acid |
| Molecular formula | C13H17N3O5 |
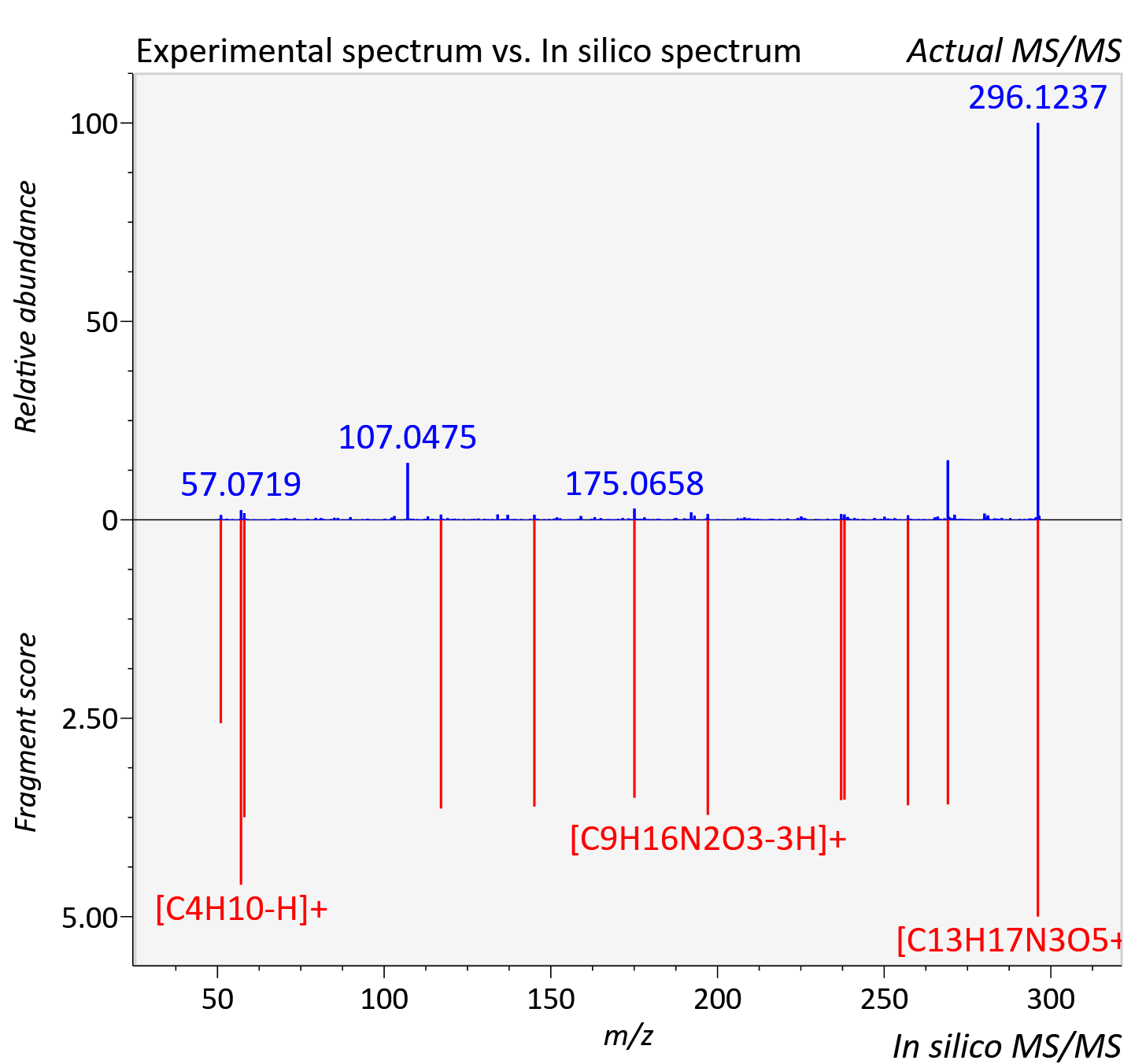
Experimental data
| Retention time | 29.92 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 296.125 | Theoretical mz | 296.124 |
| Error | 3.07 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.1083 |
Identifiers and class information
| Inchi key | NDUYBZNXWIUCHG-UHFFFAOYSA-N |
|---|---|
| Smiles | C1CCC(C1)C(=O)NC(CN2C=CC(=O)NC2=O)C(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P08473 | MME | Neprilysin | T05409 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q16478 | GRIK5 | Glutamate receptor ionotropic kainate 5 | T87930 | SEA |
| P42262 | GRIA2 | Glutamate receptor ionotropic, AMPA 2 | T42392 | SEA |
| P51681 | CCR5 | C-C chemokine receptor type 5 | T09960 | SEA |
| Q9BYF1 | ACE2 | Angiotensin-converting enzyme 2 | T27889 | SEA |
| P05556 | ITGB1 | Integrin alpha2/beta1 | T01851 | SEA |
| O00763 | ACACB | Acetyl-CoA carboxylase 2 | T08922 | SEA |
| P26010 | ITGB7 | Integrin alpha-4/beta-7 | T07403 | SEA |
| P41231 | P2RY2 | Purinergic receptor P2Y2 | T93515 | SEA |
| Q15077 | P2RY6 | Pyrimidinergic receptor P2Y6 | T90782 | SEA |
| P03951 | F11 | Coagulation factor XI | T46040 | SEA |
| O14939 | PLD2 | Phospholipase D2 | T10383 | SEA |
| Q13451 | FKBP5 | Peptidyl-prolyl cis-trans isomerase FKBP5 | T98337 | SEA |
| P33316 | DUT | dUTP pyrophosphatase | T39054 | SEA |
| P13612 | ITGA4 | Integrin alpha-4 | T97587 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| Q96SW2 | CRBN | Protein cereblon | T27861 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T05409 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P08473 | MME |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T42392 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42262 | GRIA2 |
| T09960 | DI0184 | Human immunodeficiency virus disease | [ICD-11: 1C60-1C62] | P51681 | CCR5 |
| T27889 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | Q9BYF1 | ACE2 |
| T27889 | DI0217 | Intrathoracic organs injury | [ICD-11: NB32] | Q9BYF1 | ACE2 |
| T27889 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | Q9BYF1 | ACE2 |
| T01851 | DI0060 | Brain cancer | [ICD-11: 2A00] | P05556 | ITGB1 |
| T01851 | DI0238 | Lung cancer | [ICD-11: 2C25] | P05556 | ITGB1 |
| T01851 | DI0365 | Retinopathy | [ICD-11: 9B71] | P05556 | ITGB1 |
| T08922 | DI0417 | Type 2 diabetes mellitus | [ICD-11: 5A11] | O00763 | ACACB |
| T07403 | DI0107 | Crohn disease | [ICD-11: DD70] | P26010 | ITGB7 |
| T93515 | DI0436 | Visual system disease | [ICD-11: 9E1Z] | P41231 | P2RY2 |
| T90782 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15077 | P2RY6 |
| T46040 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | P03951 | F11 |
| T46040 | DI0089 | Clotting disorder | [ICD-11: 3B4Z] | P03951 | F11 |
| T10383 | DI0091 | Coagulation defect | [ICD-11: 3B10] | O14939 | PLD2 |
| T39054 | DI0238 | Lung cancer | [ICD-11: 2C25] | P33316 | DUT |
| T39054 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P33316 | DUT |
| T97587 | DI0107 | Crohn disease | [ICD-11: DD70] | P13612 | ITGA4 |
| T97587 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P13612 | ITGA4 |
| T97587 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P13612 | ITGA4 |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T27861 | DI0060 | Brain cancer | [ICD-11: 2A00] | Q96SW2 | CRBN |
| T27861 | DI0123 | Diffuse large B-cell lymphoma | [ICD-11: 2A81] | Q96SW2 | CRBN |
| T27861 | DI0235 | Liver cancer | [ICD-11: 2C12] | Q96SW2 | CRBN |
| T27861 | DI0239 | Lupus erythematosus | [ICD-11: 4A40] | Q96SW2 | CRBN |
| T27861 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q96SW2 | CRBN |
| T27861 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | Q96SW2 | CRBN |
| T27861 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | Q96SW2 | CRBN |
| T27861 | DI0368 | Sarcoidosis | [ICD-11: 4B20] | Q96SW2 | CRBN |
| T27861 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q96SW2 | CRBN |
