Compound details
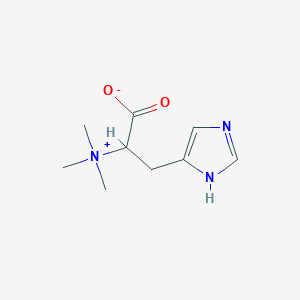
L-Histidine trimethylbetaine
| Compound ID | CDAMM00084 |
|---|---|
| Common name | L-Histidine trimethylbetaine | IUPAC name | 3-(1H-imidazol-5-yl)-2-(trimethylazaniumyl)propanoate |
| Molecular formula | C9H15N3O2 |
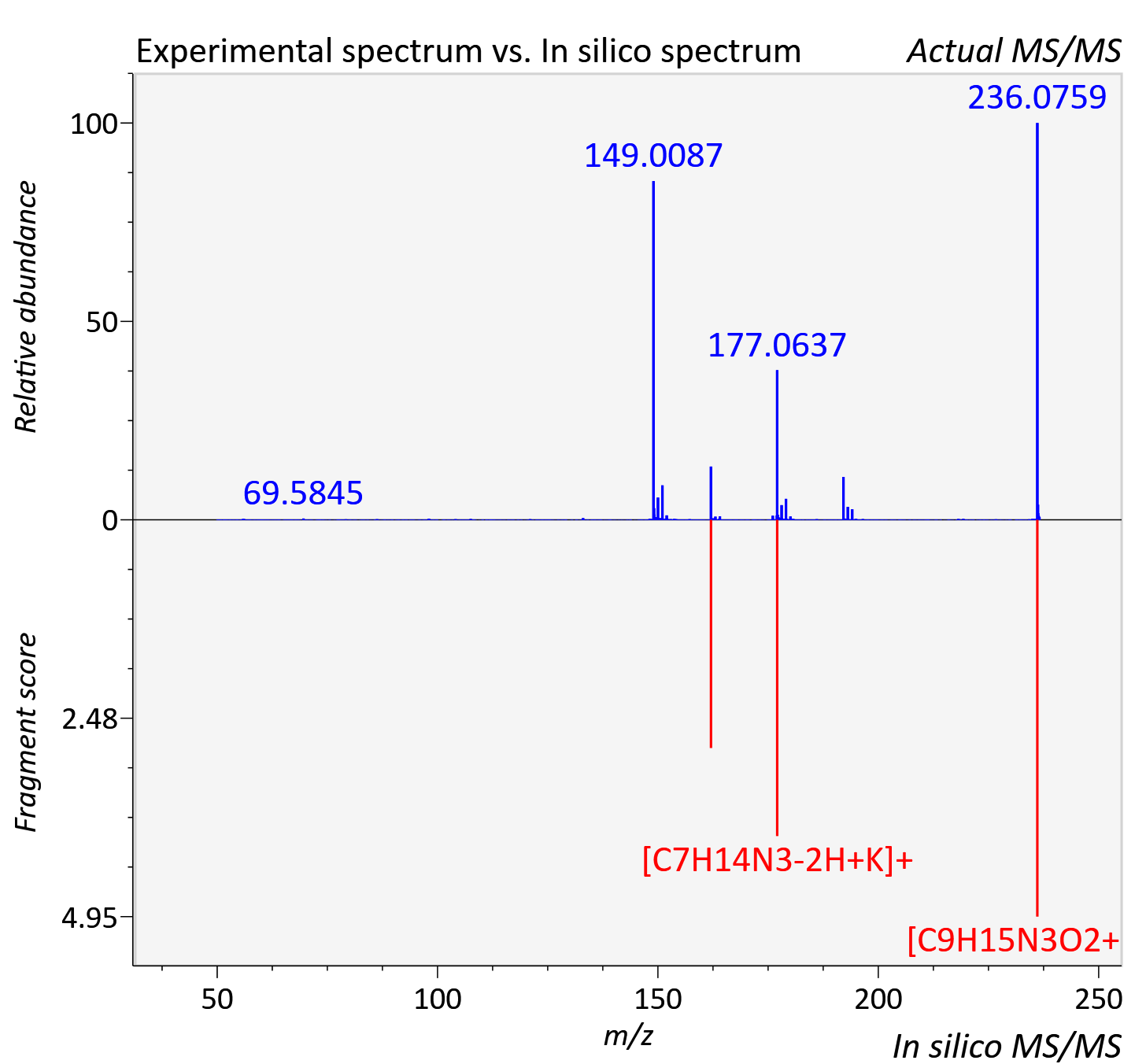
Experimental data
| Retention time | 5.93 |
|---|---|
| Adduct | [M+K]+ |
| Actual mz | 236.079 | Theoretical mz | 236.08 |
| Error | 1.18 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.707 |
Identifiers and class information
| Inchi key | GPPYTCRVKHULJH-UHFFFAOYNA-N |
|---|---|
| Smiles | C[N+](C)(C)C(CC1=CN=CN1)C(=O)[O-] |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P24046 | GABRR1 | GABA receptor rho-1 subunit | T99665 | SEA |
| Q96IY4 | CPB2 | Carboxypeptidase B2 isoform A | T98022 | SEA |
| Q9H3N8 | HRH4 | Histamine H4 receptor | T26500 | SEA |
| P15086 | CPB1 | Carboxypeptidase B | T36725 | SEA |
| Q9BYF1 | ACE2 | Angiotensin-converting enzyme 2 | T27889 | SEA |
| P28476 | GABRR2 | GABA(A) receptor rho2 | T90682 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T98022 | DI0073 | Cerebral ischaemia | [ICD-11: 8B1Z] | Q96IY4 | CPB2 |
| T98022 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q96IY4 | CPB2 |
| T98022 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | Q96IY4 | CPB2 |
| T26500 | DI0037 | Asthma | [ICD-11: CA23] | Q9H3N8 | HRH4 |
| T26500 | DI0039 | Atopic eczema | [ICD-11: EA80] | Q9H3N8 | HRH4 |
| T26500 | DI0351 | Psoriasis | [ICD-11: EA90] | Q9H3N8 | HRH4 |
| T26500 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | Q9H3N8 | HRH4 |
| T26500 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q9H3N8 | HRH4 |
| T27889 | DI0009 | Acute diabete complication | [ICD-11: 5A2Y] | Q9BYF1 | ACE2 |
| T27889 | DI0217 | Intrathoracic organs injury | [ICD-11: NB32] | Q9BYF1 | ACE2 |
| T27889 | DI0356 | Pulmonary hypertension | [ICD-11: BB01] | Q9BYF1 | ACE2 |
