Compound details
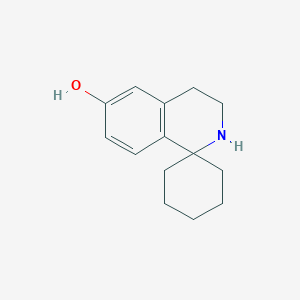
3\',4\'-dihydro-2\'H-spiro[cyclohexane-1,1\'-isoquinolin]-6\'-ol
| Compound ID | CDAMM00071 |
|---|---|
| Common name | 3\',4\'-dihydro-2\'H-spiro[cyclohexane-1,1\'-isoquinolin]-6\'-ol | IUPAC name | spiro[3,4-dihydro-2H-isoquinoline-1,1\'-cyclohexane]-6-ol |
| Molecular formula | C14H19NO |
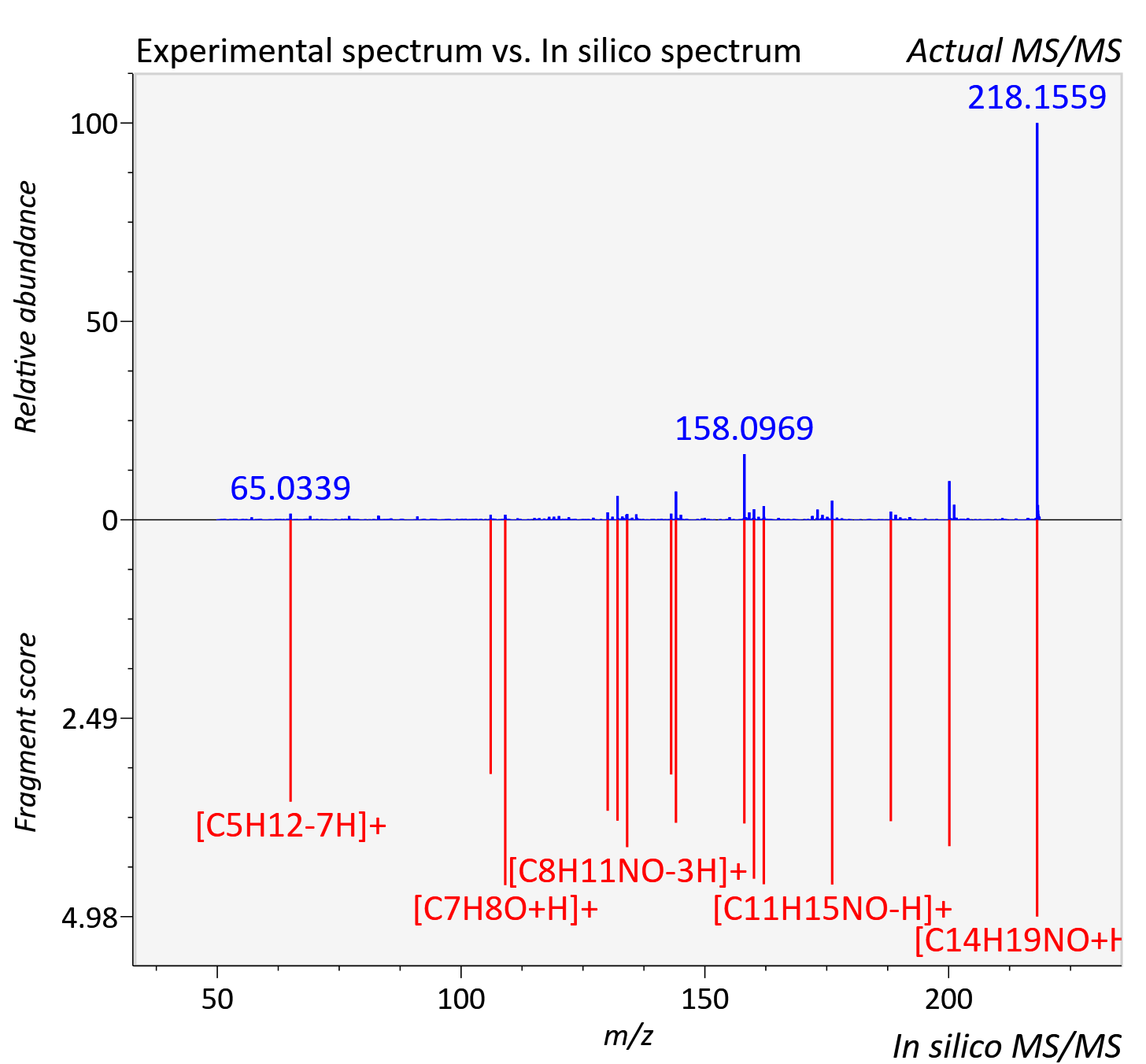
Experimental data
| Retention time | 60.54 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 218.153 | Theoretical mz | 218.154 |
| Error | 3.85 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.4803 |
Identifiers and class information
| Inchi key | ADGOCRQUBCGROE-UHFFFAOYSA-N |
|---|---|
| Smiles | C1CCC2(CC1)C3=C(CCN2)C=C(C=C3)O |
| Superclass | Organoheterocyclic compounds |
| Class | Tetrahydroisoquinolines |
Plant source
Pharmacokinetic properties
Compound-target network
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T80896 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q92731 | ESR2 |
| T80896 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | Q92731 | ESR2 |
| T80896 | DI0254 | Menopausal disorder | [ICD-11: GA30] | Q92731 | ESR2 |
| T80896 | DI0432 | Vasomotor/allergic rhinitis | [ICD-11: CA08] | Q92731 | ESR2 |
| T02506 | DI0106 | COVID-19 | [ICD-11: 1D6Y] | P03372 | ESR1 |
| T46828 | DI0022 | Allergic/hypersensitivity disorder | [ICD-11: 4A80-4A8Z] | P21918 | DRD5 |
| T58992 | DI0059 | Bowel habit change | [ICD-11: ME05] | P41143 | OPRD1 |
| T58992 | DI0218 | Irritable bowel syndrome | [ICD-11: DD91] | P41143 | OPRD1 |
| T58992 | DI0324 | Pain | [ICD-11: MG30-MG3Z] | P41143 | OPRD1 |
