Compound details
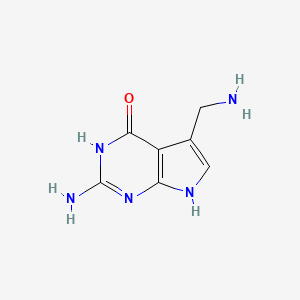
7-Aminomethyl-7-carbaguanine
| Compound ID | CDAMM00047 |
|---|---|
| Common name | 7-Aminomethyl-7-carbaguanine | IUPAC name | 2-amino-5-(aminomethyl)-3,7-dihydropyrrolo[2,3-d]pyrimidin-4-one |
| Molecular formula | C7H9N5O |
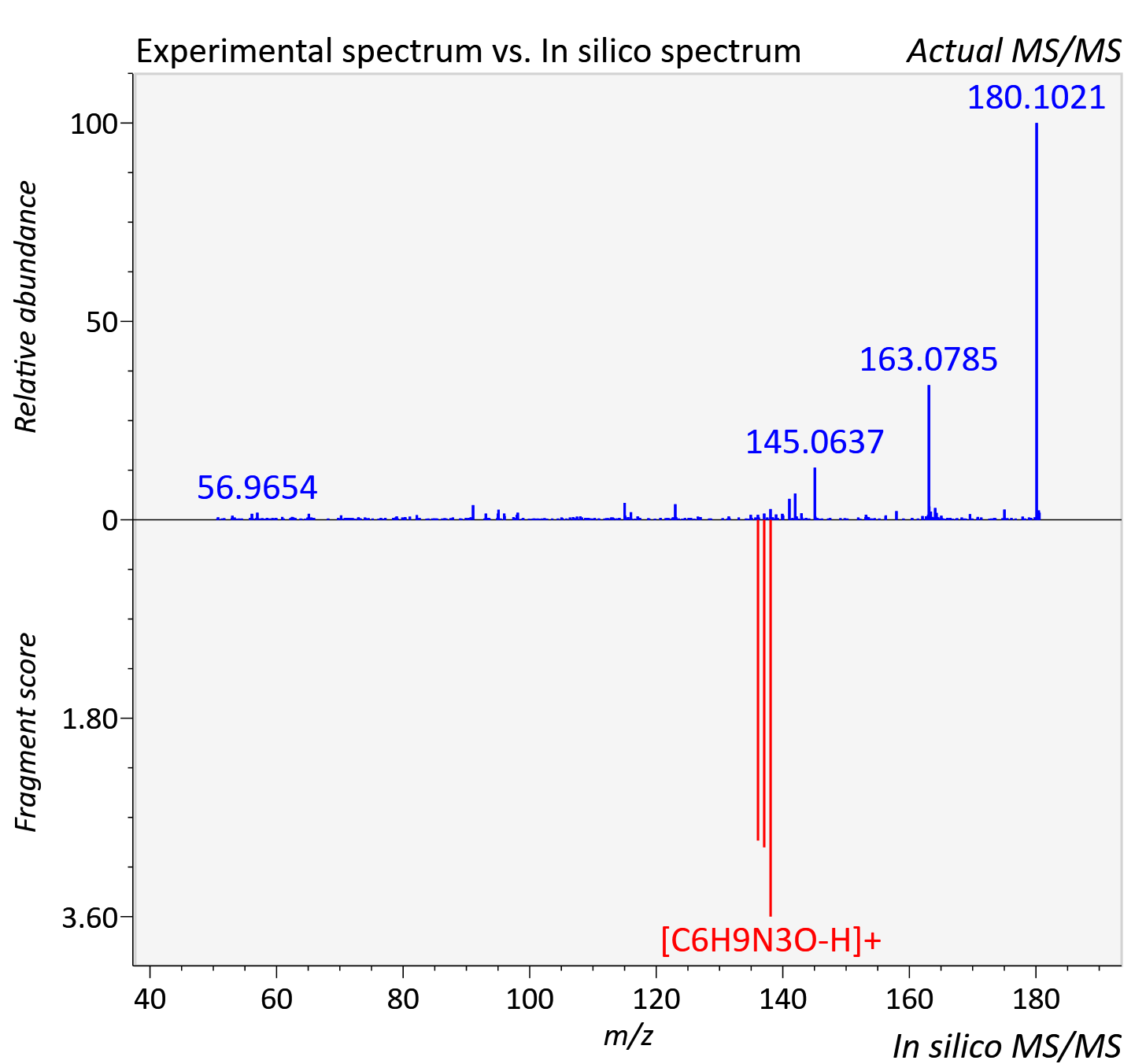
Experimental data
| Retention time | 6.5 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 180.088 | Theoretical mz | 180.088 |
| Error | 2.33 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 5.0229 |
Identifiers and class information
| Inchi key | MEYMBLGOKYDGLZ-UHFFFAOYSA-N |
|---|---|
| Smiles | C1=C(C2=C(N1)N=C(NC2=O)N)CN |
| Superclass | Organoheterocyclic compounds |
| Class | Pyrrolopyrimidines |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P04818 | TYMS | Thymidylate synthase (by homology) | T98397 | SEA |
| P00491 | PNP | Purine nucleoside phosphorylase | T78198 | SEA |
| P00374 | DHFR | Dihydrofolate reductase | T17345 | SEA |
| P47989 | XDH | Xanthine dehydrogenase | T40954 | SEA |
| P22102 | GART | GAR transformylase | T72042 | SEA |
| P15328 | FOLR1 | Folate receptor alpha | T83386 | SEA |
| P14207 | FOLR2 | Folate receptor beta | T82297 | SEA |
| Q96NT5 | SLC46A1 | Proton-coupled folate transporter | T19229 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T98397 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P04818 | TYMS |
| T98397 | DI0395 | Stomach cancer | [ICD-11: 2B72] | P04818 | TYMS |
| T78198 | DI0120 | Diabetes mellitus | [ICD-11: 5A10] | P00491 | PNP |
| T78198 | DI0167 | Gout | [ICD-11: FA25] | P00491 | PNP |
| T78198 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P00491 | PNP |
| T78198 | DI0283 | Mycosis fungoides | [ICD-11: 2B01] | P00491 | PNP |
| T78198 | DI0351 | Psoriasis | [ICD-11: EA90] | P00491 | PNP |
| T17345 | DI0207 | Indeterminate colitis | [ICD-11: DD72] | P00374 | DHFR |
| T17345 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P00374 | DHFR |
| T17345 | DI0416 | Tuberculosis | [ICD-11: 1B10-1B12] | P00374 | DHFR |
| T17345 | DI0427 | Urinary tract infection | [ICD-11: GC08] | P00374 | DHFR |
| T40954 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | P47989 | XDH |
| T40954 | DI0266 | Mineral deficiency | [ICD-11: 5B5K] | P47989 | XDH |
| T72042 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P22102 | GART |
| T83386 | DI0243 | Malaria | [ICD-11: 1F40-1F45] | P15328 | FOLR1 |
| T83386 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P15328 | FOLR1 |
| T19229 | DI0150 | Folate deficiency anaemia | [ICD-11: 3A02] | Q96NT5 | SLC46A1 |
| T19229 | DI0231 | Leukaemia | [ICD-11: 2A60-2B33] | Q96NT5 | SLC46A1 |
| T19229 | DI0306 | Nutritional deficiency | [ICD-11: 5B50-5B71] | Q96NT5 | SLC46A1 |
