Compound details
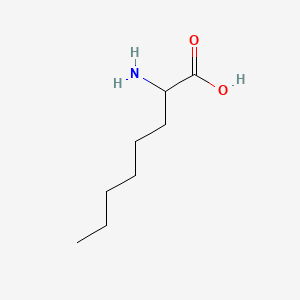
DL-2-Aminooctanoic acid
| Compound ID | CDAMM00035 |
|---|---|
| Common name | DL-2-Aminooctanoic acid | IUPAC name | 2-aminooctanoic acid |
| Molecular formula | C8H17NO2 |
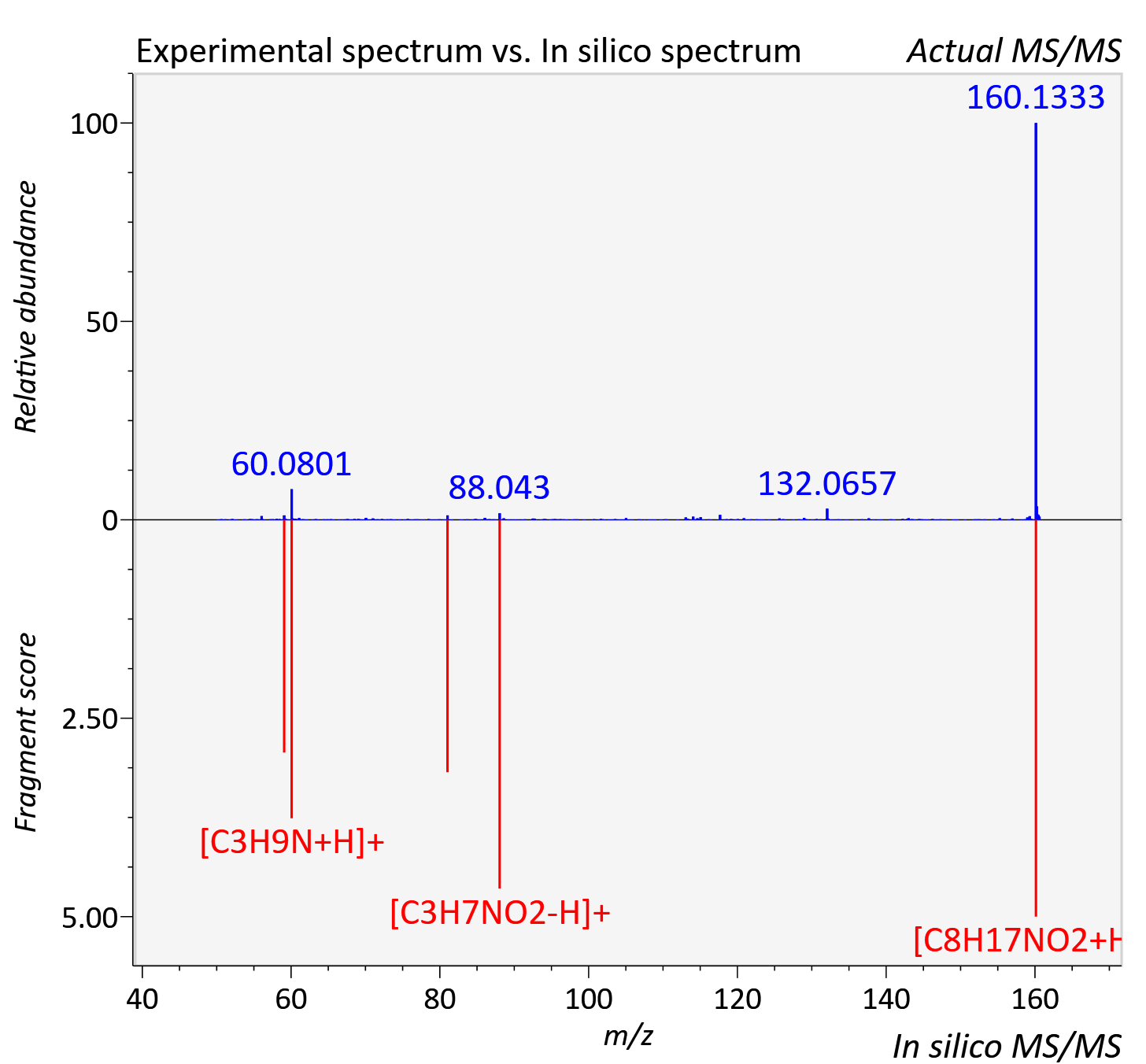
Experimental data
| Retention time | 7.37 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 160.133 | Theoretical mz | 160.133 |
| Error | 3.12 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.7502 |
Identifiers and class information
| Inchi key | AKVBCGQVQXPRLD-UHFFFAOYNA-N |
|---|---|
| Smiles | CCCCCCC(C(=O)O)N |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P07327 | ADH1A | Alcohol dehydrogenase alpha chain | T65570 | SEA |
| P29475 | NOS1 | Nitric-oxide synthase, brain | T16117 | SEA |
| P35228 | NOS2 | Nitric oxide synthase, inducible | T02703 | SEA |
| P29474 | NOS3 | Nitric-oxide synthase, endothelial | T06046 | SEA |
| P15144 | ANPEP | Aminopeptidase N | T67272 | SEA |
| Q14833 | GRM4 | Metabotropic glutamate receptor 4 | T99402 | SEA |
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| P14555 | PLA2G2A | Phospholipase A2 group IIA | T19160 | SEA |
| P43005 | SLC1A1 | Excitatory amino acid transporter 3 | T31721 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| Q13002 | GRIK2 | Glutamate receptor ionotropic kainate 2 | T58178 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| P42261 | GRIA1 | Glutamate receptor ionotropic, AMPA 1 | T33584 | SEA |
| Q16478 | GRIK5 | Glutamate receptor ionotropic kainate 5 | T87930 | SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SEA |
| P11926 | ODC1 | Ornithine decarboxylase | T60366 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SEA |
| O00519 | FAAH | Anandamide amidohydrolase | T11754 | SEA |
| Q07075 | ENPEP | Aminopeptidase A | T31956 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| P06746 | POLB | DNA polymerase beta (by homology) | T06958 | SEA |
| P08253 | MMP2 | Matrix metalloproteinase 2 | T68251 | SEA |
| P08254 | MMP3 | Matrix metalloproteinase 3 | T86702 | SEA |
| P03956 | MMP1 | Matrix metalloproteinase 1 | T52450 | SEA |
| P52732 | KIF11 | Kinesin-like protein 1 | T28484 | SEA |
| P09237 | MMP7 | Matrix metalloproteinase 7 | T73475 | SEA |
| Q9UHL4 | DPP7 | Dipeptidyl peptidase II | T50773 | SEA |
| P09211 | GSTP1 | Glutathione S-transferase Pi | T21669 | SEA |
| P08263 | GSTA1 | Glutathione S-transferase A1 | T17221 | SEA |
| Q6P179 | ERAP2 | Endoplasmic reticulum aminopeptidase 2 | T65818 | SEA |
| P00813 | ADA | Adenosine deaminase | T03661 | SEA |
| Q9ULA0 | DNPEP | Aspartyl aminopeptidase | T45287 | SEA |
| Q8TDS5 | OXER1 | Oxoeicosanoid receptor 1 | T68834 | SEA |
| Q9H228 | S1PR5 | Sphingosine 1-phosphate receptor Edg-8 | T50089 | SEA |
| P21453 | S1PR1 | Sphingosine 1-phosphate receptor Edg-1 | T13852 | SEA |
| P30307 | CDC25C | Dual specificity phosphatase Cdc25C | T40569 | SEA |
| Q9UIQ6 | LNPEP | Cystinyl aminopeptidase | T97537 | SEA |
| Q04760 | GLO1 | Glyoxalase I | T88285 | SEA |
| Q9NRA0 | SPHK2 | Sphingosine kinase 2 | T31989 | SEA |
| Q9HBW0 | LPAR2 | Lysophosphatidic acid receptor Edg-4 | T39380 | SEA |
| Q99500 | S1PR3 | Sphingosine 1-phosphate receptor Edg-3 | T11241 | SEA |
| P10826 | RARB | Retinoic acid receptor beta | T61657 | SEA |
| P53582 | METAP1 | Methionine aminopeptidase 1 | T42337 | SEA |
| P43657 | LPAR6 | Lysophosphatidic acid receptor 6 | T13484 | SEA |
| P05089 | ARG1 | Arginase-1 (by homology) | T63224 | SEA |
| O95136 | S1PR2 | Sphingosine 1-phosphate receptor Edg-5 | T47888 | SEA |
| O95977 | S1PR4 | Sphingosine 1-phosphate receptor Edg-6 | T17523 | SEA |
| Q9UPC5 | GPR34 | Probable G-protein coupled receptor 34 | T73859 | SEA |
| O00398 | P2RY10 | Putative P2Y purinoceptor 10 | T00488 | SEA |
| Q9BXC1 | GPR174 | Probable G-protein coupled receptor 174 | T87661 | SEA |
| Q9UBY5 | LPAR3 | Lysophosphatidic acid receptor Edg-7 | T95923 | SEA |
| Q92633 | LPAR1 | Lysophosphatidic acid receptor Edg-2 | T92640 | SEA |
| Q99677 | LPAR4 | Lysophosphatidic acid receptor 4 | T58130 | SEA |
| P08311 | CTSG | Cathepsin G | T86385 | SEA |
| Q9UJM8 | HAO1 | Hydroxyacid oxidase 1 | T63170 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
| P09210 | GSTA2 | Glutathione S-transferase A2 | T53510 | SEA |
| P40394 | ADH7 | Alcohol dehydrogenase class IV | T09770 | SEA |
| Q15109 | AGER | Advanced glycosylation end product receptor | T07400 | SEA |
| Q4U2R8 | SLC22A6 | Solute carrier family 22 member 6 (by homology) | T70680 | SEA |
| O60603 | TLR2 | Toll-like receptor 2 | T82078 | SEA |
| P00480 | OTC | Ornithine transcarbamylase | T52098 | SEA |
| P42263 | GRIA3 | Glutamate receptor AMPA 3 | T62391 | SEA |
| Q16348 | PEPT2 | Solute carrier family 15 member 2 | T52067 | SEA |
| Q6ZQN7 | SLCO4C1 | Organic anion transporter M1 | T36053 | SEA |
| O95749 | GGPS1 | Geranyltranstransferase | T86528 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T65570 | DI0139 | Exposure to noxious substances harmful effect | [ICD-11: NE61] | P07327 | ADH1A |
| T16117 | DI0173 | Headache | [ICD-11: 8A80-8A84] | P29475 | NOS1 |
| T16117 | DI0264 | Migraine | [ICD-11: 8A80] | P29475 | NOS1 |
| T02703 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | P35228 | NOS2 |
| T02703 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P35228 | NOS2 |
| T02703 | DI0375 | Sepsis | [ICD-11: 1G40-1G41] | P35228 | NOS2 |
| T02703 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P35228 | NOS2 |
| T06046 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P29474 | NOS3 |
| T67272 | DI0351 | Psoriasis | [ICD-11: EA90] | P15144 | ANPEP |
| T99402 | DI0331 | Parkinsonism | [ICD-11: 8A00] | Q14833 | GRM4 |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T19160 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P14555 | PLA2G2A |
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T33584 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42261 | GRIA1 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T60366 | DI0020 | African trypanosomiasis | [ICD-11: 1F51] | P11926 | ODC1 |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T11754 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | O00519 | FAAH |
| T31956 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q07075 | ENPEP |
| T06958 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P06746 | POLB |
| T68251 | DI0238 | Lung cancer | [ICD-11: 2C25] | P08253 | MMP2 |
| T86702 | DI0101 | Corneal disease | [ICD-11: 9A76-9A78] | P08254 | MMP3 |
| T86702 | DI0179 | Hepatitis virus infection | [ICD-11: 1E50-1E51] | P08254 | MMP3 |
| T86702 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | P08254 | MMP3 |
| T86702 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P08254 | MMP3 |
| T86702 | DI0287 | Myocardial infarction | [ICD-11: BA41-BA43] | P08254 | MMP3 |
| T86702 | DI0320 | Osteoarthritis | [ICD-11: FA00-FA05] | P08254 | MMP3 |
| T86702 | DI0366 | Rheumatoid arthritis | [ICD-11: FA20] | P08254 | MMP3 |
| T52450 | DI0238 | Lung cancer | [ICD-11: 2C25] | P03956 | MMP1 |
| T28484 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P52732 | KIF11 |
| T28484 | DI0172 | Head and neck cancer | [ICD-11: 2D42] | P52732 | KIF11 |
| T28484 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | P52732 | KIF11 |
| T28484 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P52732 | KIF11 |
| T28484 | DI0274 | Multiple myeloma | [ICD-11: 2A83] | P52732 | KIF11 |
| T28484 | DI0321 | Ovarian cancer | [ICD-11: 2C73] | P52732 | KIF11 |
| T28484 | DI0361 | Renal cell carcinoma | [ICD-11: 2C90] | P52732 | KIF11 |
| T28484 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P52732 | KIF11 |
| T73475 | DI0238 | Lung cancer | [ICD-11: 2C25] | P09237 | MMP7 |
| T21669 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P09211 | GSTP1 |
| T03661 | DI0016 | Adaptive immunity immunodeficiency | [ICD-11: 4A01] | P00813 | ADA |
| T03661 | DI0245 | Malignant haematopoietic neoplasm | [ICD-11: 2B33] | P00813 | ADA |
| T03661 | DI0249 | Mature B-cell leukaemia | [ICD-11: 2A82] | P00813 | ADA |
| T45287 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q9ULA0 | DNPEP |
| T50089 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | Q9H228 | S1PR5 |
| T13852 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | P21453 | S1PR1 |
| T13852 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | P21453 | S1PR1 |
| T88285 | DI0210 | Influenza | [ICD-11: 1E30-1E32] | Q04760 | GLO1 |
| T31989 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q9NRA0 | SPHK2 |
| T11241 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q99500 | S1PR3 |
| T61657 | DI0225 | Kaposi sarcoma | [ICD-11: 2B57] | P10826 | RARB |
| T63224 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P05089 | ARG1 |
| T47888 | DI0419 | Ulcerative colitis | [ICD-11: DD71] | O95136 | S1PR2 |
| T95923 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q9UBY5 | LPAR3 |
| T95923 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q9UBY5 | LPAR3 |
| T92640 | DI0146 | Fibrosis | [ICD-11: GA14-GC01] | Q92633 | LPAR1 |
| T92640 | DI0199 | Idiopathic interstitial pneumonitis | [ICD-11: CB03] | Q92633 | LPAR1 |
| T92640 | DI0351 | Psoriasis | [ICD-11: EA90] | Q92633 | LPAR1 |
| T92640 | DI0399 | Systemic sclerosis | [ICD-11: 4A42] | Q92633 | LPAR1 |
| T86385 | DI0086 | Chronic obstructive pulmonary disease | [ICD-11: CA22] | P08311 | CTSG |
| T63170 | DI0201 | Inborn carbohydrate metabolism error | [ICD-11: 5C51] | Q9UJM8 | HAO1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
| T07400 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15109 | AGER |
| T07400 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | Q15109 | AGER |
| T07400 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | Q15109 | AGER |
| T07400 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | Q15109 | AGER |
| T70680 | DI0167 | Gout | [ICD-11: FA25] | Q4U2R8 | SLC22A6 |
| T70680 | DI0206 | Inborn purine/pyrimidine/nucleotide metabolism error | [ICD-11: 5C55] | Q4U2R8 | SLC22A6 |
| T70680 | DI0310 | Ocular disease | [ICD-11: N.A.] | Q4U2R8 | SLC22A6 |
| T82078 | DI0346 | Prostate cancer | [ICD-11: 2C82] | O60603 | TLR2 |
| T52098 | DI0258 | Metabolism inborn error | [ICD-11: 5C50] | P00480 | OTC |
| T62391 | DI0298 | Neuropathy | [ICD-11: 8C0Z] | P42263 | GRIA3 |
| T36053 | DI0068 | Cardiac arrhythmia | [ICD-11: BC9Z] | Q6ZQN7 | SLCO4C1 |
| T36053 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | Q6ZQN7 | SLCO4C1 |
| T86528 | DI0057 | Bone paget disease | [ICD-11: FB85] | O95749 | GGPS1 |
| T86528 | DI0237 | Low bone mass disorder | [ICD-11: FB83] | O95749 | GGPS1 |
| T86528 | DI0267 | Mineral excesses | [ICD-11: 5B91] | O95749 | GGPS1 |
| T86528 | DI0281 | Musculoskeletal disorder | [ICD-11: FA00-FC0Z] | O95749 | GGPS1 |
