Compound details
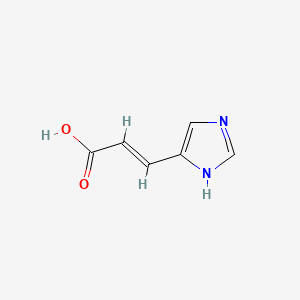
Urocanic acid
| Compound ID | CDAMM00024 |
|---|---|
| Common name | Urocanic acid | IUPAC name | (E)-3-(1H-imidazol-5-yl)prop-2-enoic acid |
| Molecular formula | C6H6N2O2 |
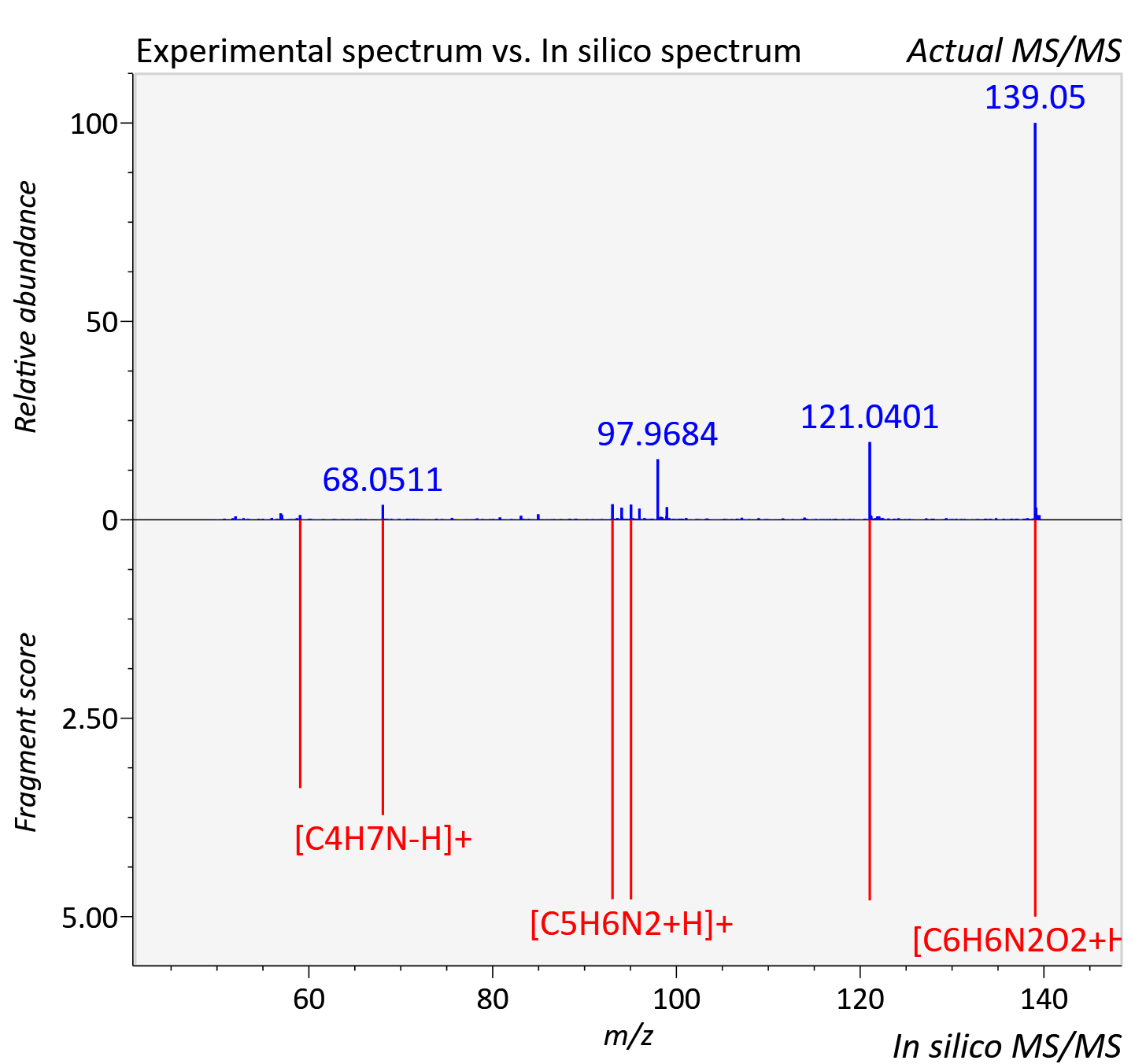
Experimental data
| Retention time | 8.68 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 139.051 | Theoretical mz | 139.05 |
| Error | 6.44 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 6.5258 |
Identifiers and class information
| Inchi key | LOIYMIARKYCTBW-OWOJBTEDSA-N |
|---|---|
| Smiles | C1=C(NC=N1)C=CC(=O)O |
| Superclass | Organoheterocyclic compounds |
| Class | Azoles |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P24046 | GABRR1 | GABA receptor rho-1 subunit | T99665 | SEA |
| P15538 | CYP11B1 | Cytochrome P450 11B1 | T84621 | SEA |
| Q96IY4 | CPB2 | Carboxypeptidase B2 isoform A | T98022 | SEA |
| P15086 | CPB1 | Carboxypeptidase B | T36725 | SEA |
| P19801 | AOC1 | Diamine oxidase | T77546 | SEA |
| P28476 | GABRR2 | GABA(A) receptor rho2 | T90682 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T84621 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | P15538 | CYP11B1 |
| T84621 | DI0108 | Cushing syndrome | [ICD-11: 5A70] | P15538 | CYP11B1 |
| T98022 | DI0073 | Cerebral ischaemia | [ICD-11: 8B1Z] | Q96IY4 | CPB2 |
| T98022 | DI0074 | Cerebral ischaemic stroke | [ICD-11: 8B11] | Q96IY4 | CPB2 |
| T98022 | DI0405 | Thrombosis | [ICD-11: DB61-GB90] | Q96IY4 | CPB2 |
