Compound details
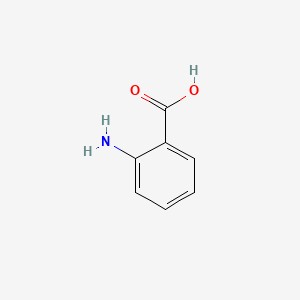
2-Aminobenzoic acid
| Compound ID | CDAMM00023 |
|---|---|
| Common name | 2-Aminobenzoic acid | IUPAC name | 2-aminobenzoic acid |
| Molecular formula | C7H7NO2 |
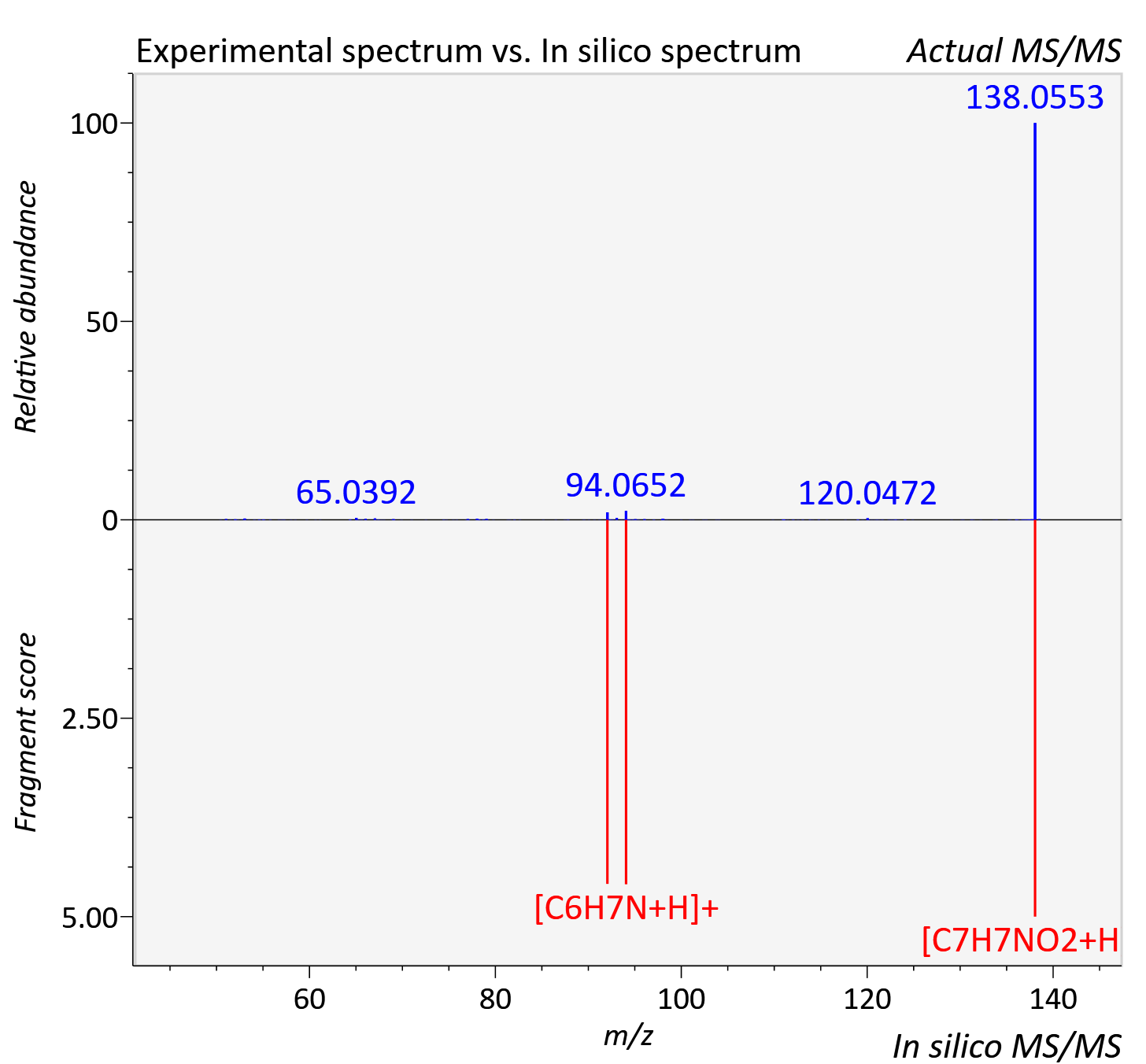
Experimental data
| Retention time | 6.56 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 138.054 | Theoretical mz | 138.055 |
| Error | 5.43 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.4784 |
Identifiers and class information
| Inchi key | RWZYAGGXGHYGMB-UHFFFAOYSA-N |
|---|---|
| Smiles | C1=CC=C(C(=C1)C(=O)O)N |
| Superclass | Benzenoids |
| Class | Benzene and substituted derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O43570 | CA12 | Carbonic anhydrase XII | T16987 | SEA |
| Q8TDS4 | HCAR2 | Hydroxycarboxylic acid receptor 2 | T88185 | SEA |
| O15379 | HDAC3 | Histone deacetylase 3 | T05090 | SEA |
| P02766 | TTR | Transthyretin | T86462 | SEA |
| P31213 | SRD5A2 | Steroid 5-alpha-reductase 2 | T71390 | SEA |
| P43166 | CA7 | Carbonic anhydrase VII | T37541 | SEA |
| P23280 | CA6 | Carbonic anhydrase VI | T06569 | SEA |
| Q9ULX7 | CA14 | Carbonic anhydrase XIV | T31992 | SEA |
| P22748 | CA4 | Carbonic anhydrase IV | T53378 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| Q92769 | HDAC2 | Histone deacetylase 2 | T51191 | SEA |
| P23141 | CES1 | Acyl coenzyme A:cholesterol acyltransferase | T76369 | SEA |
| O15554 | KCNN4 | Intermediate conductance calcium-activated potassium channel protein 4 | T42724 | SEA |
| P42330 | AKR1C3 | Aldo-keto-reductase family 1 member C3 | T60857 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| O75164 | KDM4A | Lysine-specific demethylase 4A | T18904 | SEA |
| Q6B0I6 | KDM4D | Lysine-specific demethylase 4D | T17036 | SEA |
| Q16719 | KYNU | Kynureninase | T99912 | SEA |
| P15090 | FABP4 | Fatty acid binding protein adipocyte | T07217 | SEA |
| P08575 | PTPRC | Leukocyte common antigen | T43115 | SEA |
| Q9Y4C1 | KDM3A | Lysine-specific demethylase 3A | T25362 | SEA |
| O15054 | KDM6B | Lysine-specific demethylase 6B | T56057 | SEA |
| P27695 | APEX1 | DNA-(apurinic or apyrimidinic site) lyase | T13348 | SEA |
| P08263 | GSTA1 | Glutathione S-transferase A1 | T17221 | SEA |
| Q99523 | SORT1 | Neurotensin receptor 3 | T63233 | SEA |
| Q9ULW8 | PADI3 | Protein-arginine deiminase type-3 | T36612 | SEA |
| Q9UM07 | PADI4 | Protein-arginine deiminase type-4 | T50429 | SEA |
| Q02127 | DHODH | Dihydroorotate dehydrogenase | T61400 | SEA |
| P42224 | STAT1 | Signal transducer and activator of transcription 1-alpha/beta | T64205 | SEA |
| P22736 | NR4A1 | Nuclear receptor subfamily 4 group A member 1 | T41119 | SEA |
| Q9Y256 | RCE1 | Prenyl protein specific protease | T10756 | SEA |
| Q9UPY5 | SLC7A11 | Cystine/glutamate transporter | T11615 | SEA |
| Q15109 | AGER | Advanced glycosylation end product receptor | T07400 | SEA |
| Q04609 | FOLH1 | Glutamate carboxypeptidase II | T97071 | SEA |
| P09172 | DBH | Dopamine beta hydroxylase | T74937 | SEA |
| P46952 | HAAO | 3-hydroxyanthranilate oxygenase | T46527 | SEA |
| P09936 | UCHL1 | Ubiquitin thioesterase L1 | T56051 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T16987 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | O43570 | CA12 |
| T16987 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | O43570 | CA12 |
| T88185 | DI0203 | Inborn lipid metabolism error | [ICD-11: 5C52] | Q8TDS4 | HCAR2 |
| T05090 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | O15379 | HDAC3 |
| T05090 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | O15379 | HDAC3 |
| T86462 | DI0026 | Amyloidosis | [ICD-11: 5D00] | P02766 | TTR |
| T71390 | DI0348 | Prostate hyperplasia | [ICD-11: GA90] | P31213 | SRD5A2 |
| T06569 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P23280 | CA6 |
| T06569 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P23280 | CA6 |
| T31992 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q9ULX7 | CA14 |
| T31992 | DI0204 | Inborn metabolism deficiency | [ICD-11: 5C50-5C59] | Q9ULX7 | CA14 |
| T53378 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | P22748 | CA4 |
| T53378 | DI0166 | Glaucoma | [ICD-11: 9C61] | P22748 | CA4 |
| T53378 | DI0372 | Seborrhoeic dermatitis | [ICD-11: EA81] | P22748 | CA4 |
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T51191 | DI0241 | Lymphoma | [ICD-11: 2A80-2A86] | Q92769 | HDAC2 |
| T51191 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | Q92769 | HDAC2 |
| T76369 | DI0335 | Peroxisomal disease | [ICD-11: 5C57] | P23141 | CES1 |
| T76369 | DI0398 | Synthesis disorder | [ICD-11: 5C52-5C59] | P23141 | CES1 |
| T42724 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | O15554 | KCNN4 |
| T42724 | DI0087 | Chronic pain | [ICD-11: MG30] | O15554 | KCNN4 |
| T60857 | DI0145 | Female pelvic pain | [ICD-11: GA34] | P42330 | AKR1C3 |
| T99912 | DI0046 | Bacterial infection | [ICD-11: 1A00-1C4Z] | Q16719 | KYNU |
| T99912 | DI0062 | Breast cancer | [ICD-11: 2C60-2C6Y] | Q16719 | KYNU |
| T43115 | DI0012 | Acute myeloid leukaemia | [ICD-11: 2A60] | P08575 | PTPRC |
| T43115 | DI0414 | Transplanted organ/tissue | [ICD-11: QB63] | P08575 | PTPRC |
| T13348 | DI0365 | Retinopathy | [ICD-11: 9B71] | P27695 | APEX1 |
| T13348 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P27695 | APEX1 |
| T63233 | DI0153 | Frontotemporal dementia | [ICD-11: 6D83] | Q99523 | SORT1 |
| T61400 | DI0188 | Hyper-lipoproteinaemia | [ICD-11: 5C80] | Q02127 | DHODH |
| T61400 | DI0275 | Multiple sclerosis | [ICD-11: 8A40] | Q02127 | DHODH |
| T64205 | DI0383 | Skin and skin-structure infection | [ICD-11: 1F28-1G0Z] | P42224 | STAT1 |
| T11615 | DI0054 | Body-focused behaviour disorder | [ICD-11: 6B25] | Q9UPY5 | SLC7A11 |
| T07400 | DI0025 | Alzheimer disease | [ICD-11: 8A20] | Q15109 | AGER |
| T07400 | DI0083 | Chronic kidney disease | [ICD-11: GB61] | Q15109 | AGER |
| T07400 | DI0115 | Dementia | [ICD-11: 6D80-6D8Z] | Q15109 | AGER |
| T07400 | DI0196 | Hypotension | [ICD-11: BA20-BA21] | Q15109 | AGER |
| T97071 | DI0122 | Diagnostic imaging | [ICD-11: N.A.] | Q04609 | FOLH1 |
| T97071 | DI0346 | Prostate cancer | [ICD-11: 2C82] | Q04609 | FOLH1 |
| T74937 | DI0175 | Heart failure | [ICD-11: BD10-BD1Z] | P09172 | DBH |
| T74937 | DI0341 | Post-traumatic stress disorder | [ICD-11: 6B40] | P09172 | DBH |
| T56051 | DI0391 | Solid tumour/cancer | [ICD-11: 2A00-2F9Z] | P09936 | UCHL1 |
