Compound details
L-Valine
| Compound ID | CDAMM00010 |
|---|---|
| Common name | L-Valine | IUPAC name | (2S)-2-amino-3-methylbutanoic acid |
| Molecular formula | C5H11NO2 |
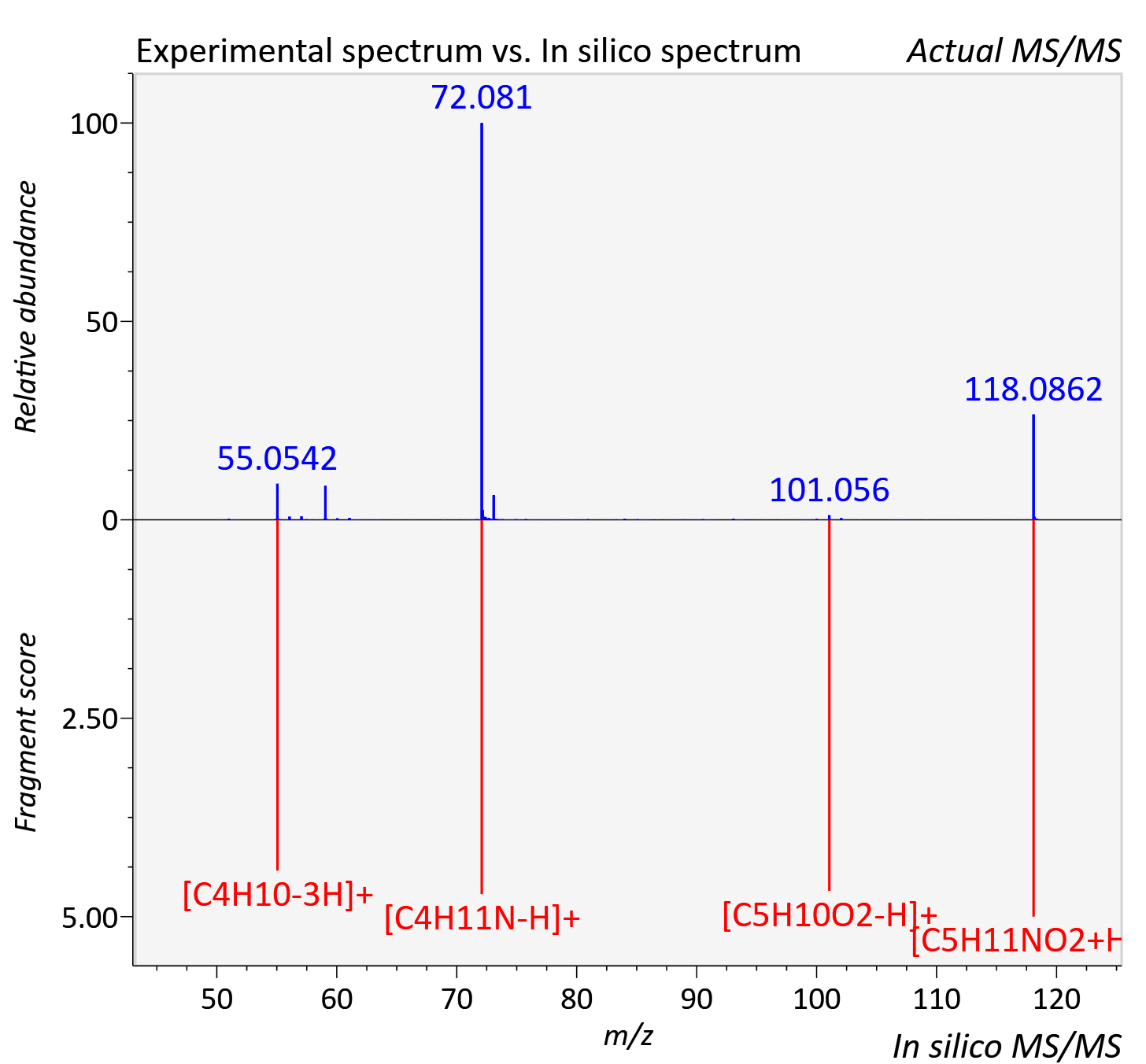
Experimental data
| Retention time | 7.25 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 118.087 | Theoretical mz | 118.086 |
| Error | 3.81 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.0663 |
Identifiers and class information
| Inchi key | KZSNJWFQEVHDMF-SGAVLPGINA-N |
|---|---|
| Smiles | CC(C)C(C(=O)O)N |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| O00222 | GRM8 | Metabotropic glutamate receptor 8 | T77548 | SEA |
| P43005 | SLC1A1 | Excitatory amino acid transporter 3 | T31721 | SEA |
| P39086 | GRIK1 | Glutamate receptor ionotropic kainate 1 | T73495 | SEA |
| Q13002 | GRIK2 | Glutamate receptor ionotropic kainate 2 | T58178 | SEA |
| O15303 | GRM6 | Metabotropic glutamate receptor 6 | T55956 | SEA |
| Q16478 | GRIK5 | Glutamate receptor ionotropic kainate 5 | T87930 | SEA |
| Q13003 | GRIK3 | Glutamate receptor ionotropic kainate 3 | T68876 | SEA |
| Q9H4A4 | RNPEP | Aminopeptidase B | T57818 | SEA |
| Q07075 | ENPEP | Aminopeptidase A | T31956 | SEA |
| P43003 | SLC1A3 | Excitatory amino acid transporter 1 | T86582 | SEA |
| Q9Y3Q0 | NAALAD2 | NAALADase II | T70036 | SEA |
| Q9UPY5 | SLC7A11 | Cystine/glutamate transporter | T11615 | SEA |
Target associated diseases
| TTD_ID | Disease_ID | Disease name | ICD_11 | Uniprot ID | Gene names |
|---|---|---|---|---|---|
| T73495 | DI0134 | Epilepsy/seizure | [ICD-11: 8A61-8A6Z] | P39086 | GRIK1 |
| T73495 | DI0396 | Substance abuse | [ICD-11: 6C40] | P39086 | GRIK1 |
| T31956 | DI0190 | Hypertension | [ICD-11: BA00-BA04] | Q07075 | ENPEP |
| T11615 | DI0054 | Body-focused behaviour disorder | [ICD-11: 6B25] | Q9UPY5 | SLC7A11 |
