Compound details
L-Proline
| Compound ID | CDAMM00009 |
|---|---|
| Common name | L-Proline | IUPAC name | (2S)-pyrrolidine-2-carboxylic acid |
| Molecular formula | C5H9NO2 |
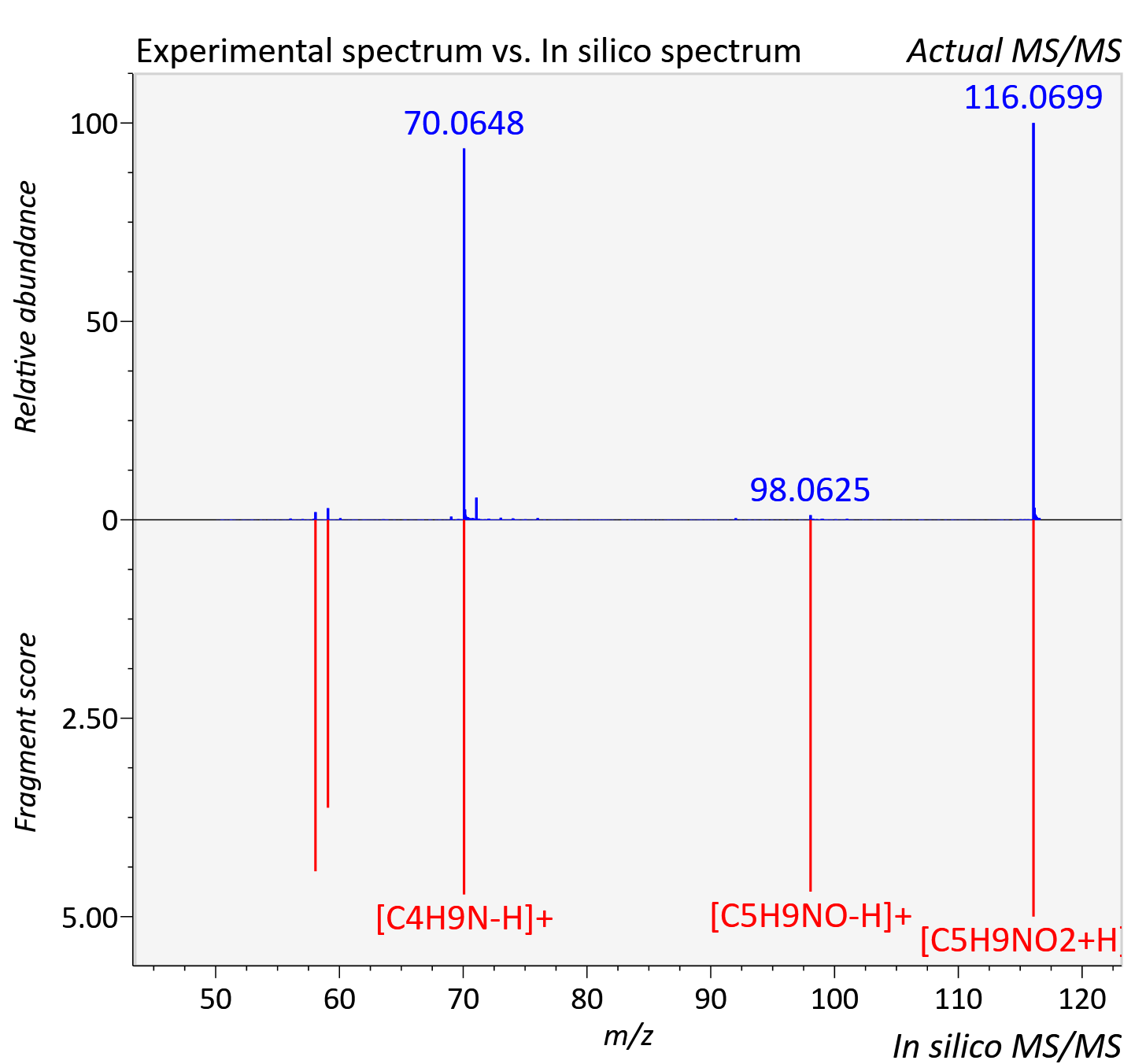
Experimental data
| Retention time | 6.62 |
|---|---|
| Adduct | [M+H]+ |
| Actual mz | 116.071 | Theoretical mz | 116.071 |
| Error | 3.44 |
| Ionizaton mode | Positive |
| Instrument type | LC-MS/MS-QTOF, spectrum predicted by MS-DIAL v4.9 integrated with MS-FINDER 3.60 |
| Score | 8.4175 |
Identifiers and class information
| Inchi key | ONIBWKKTOPOVIA-BYPYZUCNSA-N |
|---|---|
| Smiles | C1CC(NC1)C(=O)O |
| Superclass | Organic acids and derivatives |
| Class | Carboxylic acids and derivatives |
Plant source
Pharmacokinetic properties
Compound-target network
Protein targets associated with phytocompound
| Uniprot ID | Gene name | Target name | TTD_ID | Prediction source |
|---|---|---|---|---|
| P12955 | PEPD | Xaa-Pro dipeptidase | T94823 | SEA |
| P15169 | CPN1 | Carboxypeptidase N, catalytic subunit | T34471 | SEA |
| Q9NY46 | SCN3A | Sodium channel protein type III alpha subunit | T76937 | SEA |
| P27708 | CAD | Aspartate carbamoyltransferase | T24548 | SEA |
